INTRODUCTION
We previously identified early growth response 1 (Egr1) expression in the pregnant mouse uterus using the microarray technique (Cheon et al., 2002). It is known that Egr1 is an immediate-early gene like c-Fos, and a non-housekeeping gene. Its expression is not ubiquitous in various tissues such as tendon, cartilage, bone, skeletal muscle, dermis, and stroma in the embryo. In developing tendons, Egr1 expression is not detected just in the myotendinous junction but around long tendons in the mouse embryo (McMahon et al., 1990; Lejard et al., 2011). In the adult stage, Egr1 expression is detected in many tissues, including the cortex, adipose tissue, mammary gland, ovary, uterus, and thymus, but its expression is not ubiquitous (Sukhatme et al., 1988; Guo et al., 2014; Milet et al., 2017).
The Egr1 gene is rapidly and transiently activated in most of the cell types by various stimulations, including mitogens. It was identified as nerve growth factor induced-A (NGFI-A) in the late 1980s. This gene and protein have had many names. In 1987, Milbrandt reported the NGFI-A gene, which was rapidly and strongly induced by nerve growth factor (NGF) in PC12 cells. He showed that this gene encodes a transcription factor with zinc finger domains (Milbrandt, 1987). In 1988, Christy and colleagues identified this gene in mouse 3T3 fibroblasts after serum treatment and named it Zif268 due to its zinc-finger structure and rapid expression (Christy et al., 1988). Sukhatme et al. (1988) cloned this gene in NIH3T3 cells after 12-O-tetradecanoylphorbol-13-acetate (TPA) administration and named it TPA inducible sequence 8 (TIS8). Lemaire et al. (1988) cloned this gene in serum-treated NIH 3T3 and F9 cells and named it Krox-24 (Krüppel box 24) due to its Krüppel-like zinc finger structure. Christy et al. (1988) identified this gene in 3T3 cells after the administration of serum growth factor and named it zif268, in reference to its three tandem zinc finger sequences. In addition, a few names were also found (Mello et al., 1992; NCBI, 2025). For example, it was named G0 to G1 switch gene 30 (GOS30) because it was activated during the G0/G1 transition. Now, it’s officially called Egr1, which stands for Egr1, as it turns on quickly in response to various stimuli, including growth factors, stress, or damage.
The immediate early gene means that its expression doesn’t need new proteins to be made in order to turn on, and it acts as a transcription factor to help regulate other genes right away. The Egr1 gene is highly conserved across invertebrates and mammals, including mice and humans (Burmeister & Fernald, 2005; Ugajin et al., 2016). The Egr1 product belongs to the early growth response (EGR) protein family, Cys2His2 (C2H2) class zinc-finger (ZF) nuclear proteins. This transcription family includes EGR1, EGR2, EGR3, and EGR4, and is involved in such cell differentiation, proliferation, and stress responses (O’Donovan et al., 1999). In the case of Egr1, there are three ZF encoding sequences, and the molecular weight of EGR1 is approximately 59–60 kDa, primarily localized in the nucleus.
The biological roles of the Egr1 protein have been evaluated through studies by many groups. Functionally, EGR1 plays crucial roles in various biological processes, including developmental processes such as cell proliferation, differentiation, inflammation, and cell death. Moreover, EGR1 can function as either a transcriptional activator or repressor depending on the cellular context and can work as a master switch of divergent gene families (Yan et al., 2000; Baek et al., 2022). For instance, in macrophages, EGR1 suppresses inflammatory enhancer activity by recruiting the NuRD corepressor complex, acting as a brake on chronic inflammation (Trizzino et al., 2021). On the other hand, in cancer cells such as those of the prostate, EGR1 can paradoxically promote angiogenesis and metastasis through upregulation of pro-tumorigenic factors (Li et al., 2019). It is known as a cancer-suppressing or promoting gene, and is also required for differentiation and mitogenesis. In this review, we briefly summarize current knowledge of EGR1 and its gene anatomy. Here, the nomenclature of genes and proteins primarily follows the mouse system.
The Egr1 is a protein coding gene with aliases TIS8, AT225, GOS30, NGFI-A, ZNF225, KROX-24, zinc finger protein identified at 268 nm (ZIF268), and ZIF-268 in human and ETR103, TIS8, Zenk, Egr-1, NGFIA, Zfp-6, Kron-1, Krox24, Krox-24, NGF1-A, NGFI-A, Zif268, and A530045N19Rik in mouse (NCBI, 2025; Dyer et al., 2025). This gene is localized on chromosome 5 (138,465,479–138,469,303 forward strand; GRCh38:CM000667.2; band 5q31.2) and chromosome 18 (34,994,260-34,998,009 forward strand; GRCm39:CM001011.3; band 18B1) in humans and mice, respectively (Fig. 1). The Egr-1 gene spans about 3.8 kb and 4.4 kb in both human and mouse, respectively. Both in human and mouse, there are two exons and one intron, and 5’ flanking sequence (Tsai-Morris et al., 1988).
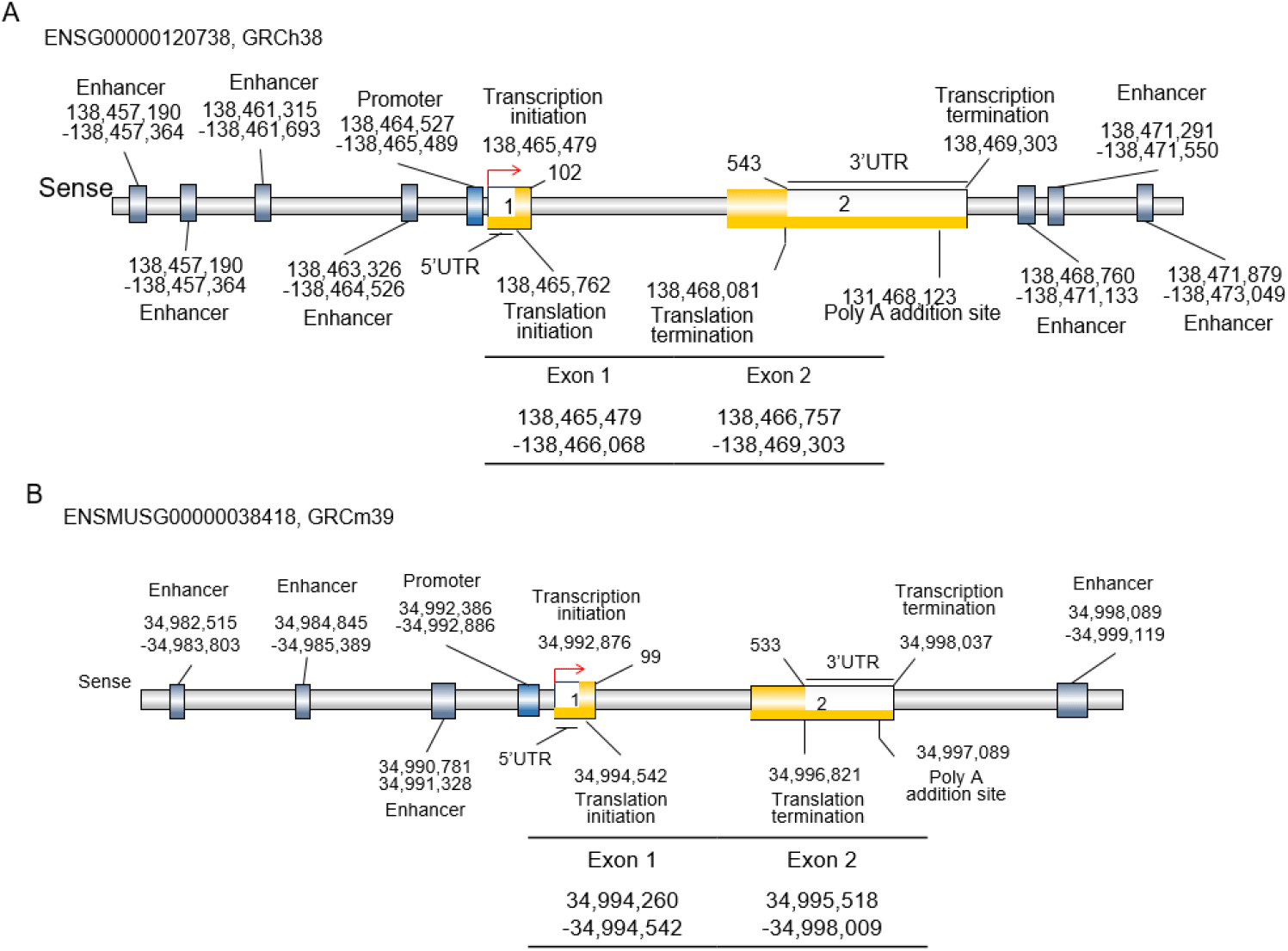
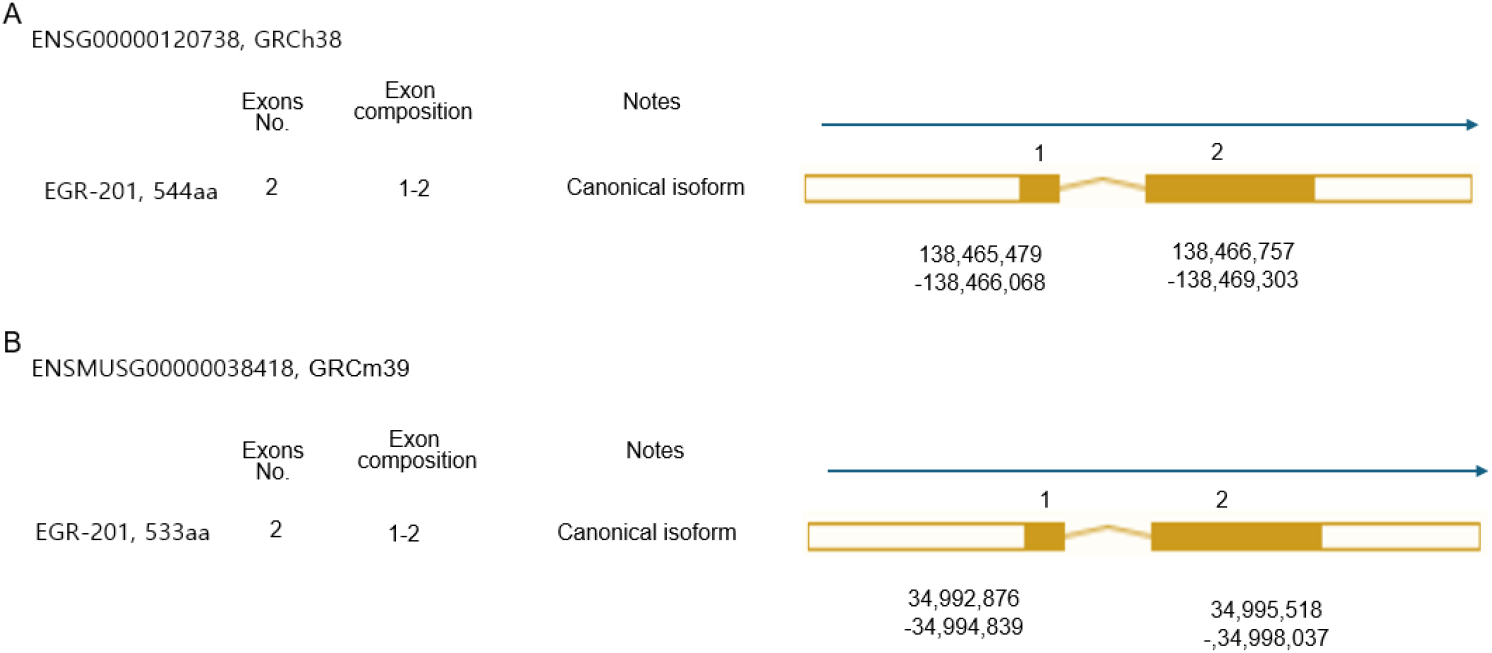
Exon 1 includes the 5’ untranslated region (UTR), transcription initiation sequence, and translation initiation codon. Exon 2 includes the stop codon, the 3’UTR, one PolyA signal sequence and one PolyA site and PolyA signal sequence in both human and mouse (Figs. 1 and 2). The exons encode a 544 amino acid precursor protein in humans and 533 amino acids in mice (Figs. 2 and 3). Egr1 is well conserved in vertebrates (Makałowski et al., 1996; Solari et al., 1999; Burmeister & Fernald, 2005), and the similarity between human and mouse is approximately 86.42% in amino acid sequences and 85% identity in protein-coding sequences (Fig. 2B and C).

The mouse Egr1 gene is constructed with a single promoter and a promoter flank. In the core promoter of Egr1 of mouse, the TATA and CCAAT boxes are identified, and their sequences are located at –26 and –337, respectively (Tsai-Morris et al., 1988). Interestingly, on the other hand, in the case of human, there are two promoters. One of them includes the promoter flank (located at 138,458,518–138,459,018), and the other includes the core promoter in front of exon 1 (located at 138,464,527–138,465,489). A TATA box was identified at –26 (Schwachtgen et al., 2000), but no CCAAT was identified in full promoter sequencing (Fig. 2A and B). Because TATA-box and CAT-box are thought to be absent in house-keeping genes, including protooncogenes (Huang et al., 1997b; Zhu et al., 2008). The core promoter is located upstream of exon 1. At the promoter, many conserved regulatory elements, including c-Fos binding sequence, activator protein 1 (AP1), potential cAMP response elements (CRE), hypoxia response element (HRE), EGR1 binding site (EBS), specificity protein 1 (SP1) response elements (Sp1RE), nuclear factor kappa-light-chain-enhancer of activated B cells (NFkB) site, and serum response elements (SREs) are identified (Tsai-Morris et al., 1988; Havis & Duprez, 2020). For example, the serum response factor binding motif, CArG box (CC(A/T)6GG) is depicted in Fig. 2. Those are summarized in Table 1.
EGR1, early growth response 1.
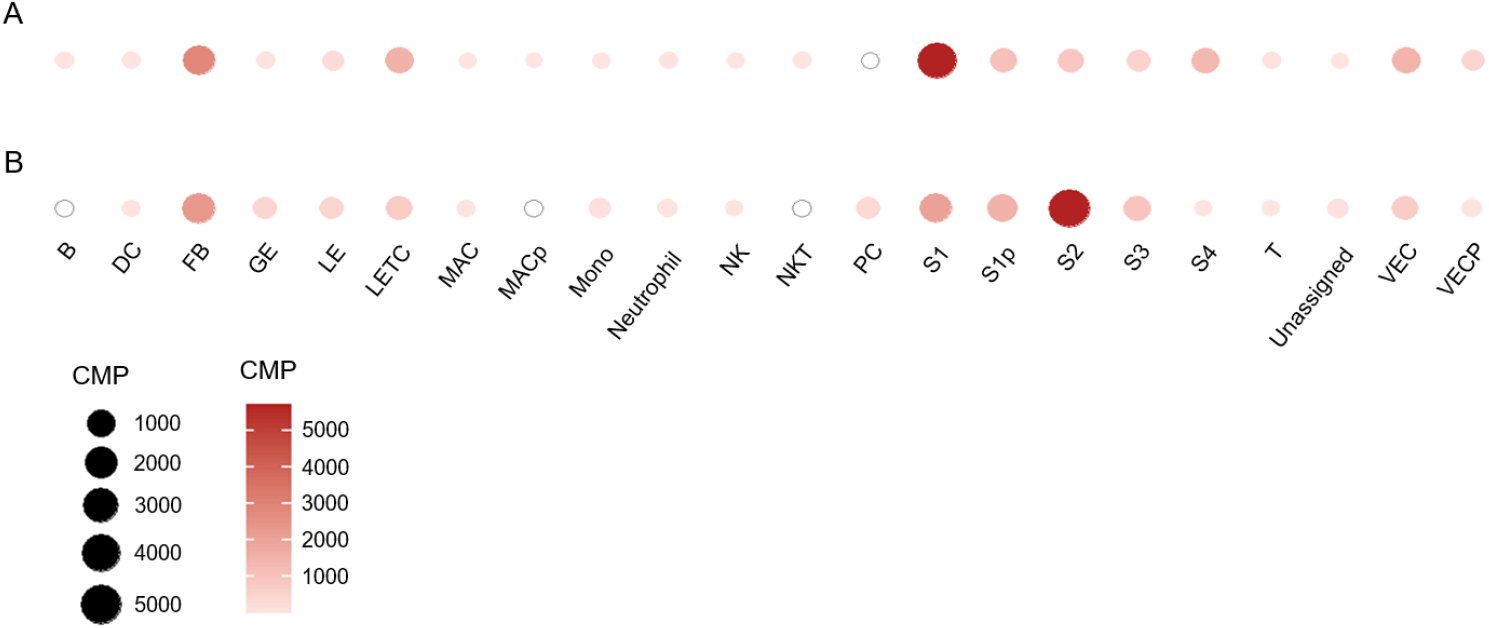
Enhancers are distal cis-elements believed to control gene expression from core promoters in a developmental stage and tissue-specific manner (Spitz & Furlong, 2012; Ray-Jones et al., 2025). However, tissue specificity is low, with a 0.22 tau index (The Human Protein Atlas, 2025), because its expression depends on exogenous stimuli. As seen in Fig. 1, the human EGR1 gene has several enhancers. In humans, a proximal enhancer is localized at 138,463,326–138,464,526. As distal enhancer, there are six (3 are front of promoter and 3 are behind the exon 2) at 138,461,315– 138,461,693 (about –3.2 kb), 138,460,131–138,460,526 (about –4.4 kb), 138,457,190–138,457,364 (about –7.3 kb), 138,468,760–138,471,133 (about +4.2 kb), 138,471,291–138,471,550 (about +6.8 kb), and 138,471,879–138,473,049 (about 7.4 kb) (Fig. 1). In mouse, the Egr1 gene has four enhancers at 34,990,781–34,991,328 (about –2.1 kb), 34,984,845–34,985,389 (about –8.0 kb), 34,982,515–34,983,803 (about –10.4 kb), and 34,998,089–34,999,119 (about +5.2 kb). Humans have more distal EGR1 enhancers than mice, reflecting greater tissue specialization and more fine-tuned regulation of disease-associated genes. Interestingly, our studies showed that there are cell-type specificities in Egr1 expression by the physiological status. Egr1 mRNA is detected in various cell types, including B-cells, proliferating macrophages, and natural killer cells on day 4 of pregnancy. However, the expression of this gene in these cells is not detected on day 7 of pregnancy (Fig. 4). On the other hand, in the case of pericytes, Egr1 mRNA was not detected on day 4 but not day 7 of pregnancy. Among the stromal cells, S1 showed the highest level at day 4 and shifted to S3 by decidualization (Fig. 4). It may be related to the gene’s anatomical characters and cell proliferation.
Immediate early gene means that it is rapidly induced within minutes to hours in response to extracellular and internal stimuli, similar to Fos (Thiel & Cibelli, 2002; Havis & Duprez, 2020). In normal conditions, EGR1 displays c-Fos-like induction kinetics in various cell types, such as fibroblasts, epithelial cells, and lymphocytes. The physiological or physical inducer for EGR1 expression are numerous including stress signaling (such as glucose increase, hypoxia, UV irradiation and mechanical stimulation), endo-/para-/auto-/juxta-crines (such as hormones, growth factors, cytokines, interleukins), serum, mechanical signals, depolarization of membrane and internal signals (such as DNA damage, ischemia) (Sheng & Greenberg, 1990; Hasan et al., 2003; Chang et al., 2008; Havis & Duprez, 2020; Herchenhan et al., 2020). While the sequences of EGR1 and responses to the inducers are highly conserved between human and mouse, the expression patterns and regulatory mechanisms are different in specific contexts, such as diet-response and certain diseases (Huang et al., 1997a; Weng et al., 2012).
The external signals for Egr1 expression are mediated by cellular signal transductions (Tables 2–5). For Egr1 expression, RNA pol II recruitment depends on signal-activated transcription factors and off condition as the default condition (Gillies et al., 2017; Zhang et al., 2017). Such as mitogen-activated protein kinase (MAPK) signaling pathways through SREs on Egr1 gene for its expression (Schwachtgen et al., 2000; Bhattacharyya et al., 2008; Hoffmann et al., 2008; Rockel et al., 2009; Yang et al., 2016; Geng et al., 2019). In addition, cAMP/PKA/CREB pathway mediates Egr1 activation for hormones or cytokines (Sheng et al., 1990; Kang et al., 2007), PI3K/Akt pathway for glucose uptake and glycogenesis and the ERK1/2 MAPK pathway for decreased insulin (Franke et al., 1995; Biddinger & Kahn, 2006), MAPK/MEK/ERK/ELK1 pathway for amino acid limitation (Shan et al., 2014), cytoplasmic Ca2+ for AP1, CRE, and SER-mediated activation (Müller et al., 2012), and AMP-activated protein kinase (AMPK) for high insulin and glucose levels (Wu et al., 2017). Integrin/EGFR cross-talk is required for Egr1 expression through activation of Erk1/2 and PI3K/Akt/Forkhead pathways (Weng et al., 2012).
| Signal category | Stimulus / condition | Upstream pathway | Key TFs / elements | References |
|---|---|---|---|---|
| Basal regulation | Default OFF state | RNA Pol II recruitment requires signal-activated TFs | - |
Gillies et al., 2017; Zhang et al., 2017 |
| Mitogen / growth factor | MAPK activation | MAPK → ERK → ELK1/SRF | SREs |
Schwachtgen et al., 2000; Bhattacharyya et al., 2008; Hoffmann et al., 2008; Rockel et al., 2009; Yang et al., 2016; Geng et al., 2019 |
| Hormones / cytokines | cAMP signaling | cAMP → PKA → CREB | CRE |
Sheng et al., 1990; Kang et al., 2007 |
| Glucose / insulin | Metabolic signaling | PI3K/Akt; ERK1/2 MAPK | - |
Franke et al., 1995; Biddinger & Kahn, 2006 |
| Amino acid limitation | Nutrient stress | MAPK → MEK → ERK → ELK1 | ELK1 | Shan et al., 2014 |
| Ca2+ signaling | Cytoplasmic Ca2+ elevation | Ca2+ → AP-1 / CRE / SER | AP-1, CRE, SER | Müller et al., 2012 |
| Energy stress | High insulin / glucose | AMPK pathway | - | Wu et al., 2017 |
| ECM–growth factor cross-talk | Integrin / EGFR interaction | ERK1/2; PI3K/Akt/Forkhead | - | Weng et al., 2012 |
Egr1, early growth response 1; TF, transcription factor; MAPK, mitogen-activated protein kinase; SREs, serum response elements; CRE; cAMP response elements.
| Disease context | Upstream pathway | Key factors | References |
|---|---|---|---|
| ER stress | SRC → RAS → RAF → MEK → ERK | SRF, ELK1 | Shan et al., 2019 |
| Cancer | RTK/MAPK/ERK/ELK1; PI3K/AKT; JNK/p38; FAK/RhoA/YAP-TAZ | ELK1, AP-1, YAP/TAZ | Gitenay & Baron, 2009 |
| Osteoarthritis | FAK/Src/MAPK/ELK1/SRF; MAPK/AP-1/ELK1; β-catenin | SRF, AP-1 |
Rockel et al., 2009
Sun et al., 2019 |
| Cardiovascular disease | FAK/Src/MAPK(ERK)/ELK1/SRF; PKC/ROS/MAPK; ELK/SRF | ELK1, SRF | Khachigian, 2023 |
| Neurodegenerative / Inflammatory diseases | Ca2+/MAPK(ERK1/2, p38)/ELK1/SRF; MAPK(ERK, JNK, p38)/AP-1/ELK1; Ca2+/CaMKII/ERK/CREB/ELK1; MyD88/MAPK | ELK1, AP-1, CREB |
Bouallegue et al., 2013; Khachigian, 2023 |
Egr1, early growth response 1; MAPK, mitogen-activated protein kinase; ER, endoplasmic reticulum.
| Mechanism | Description | Context | References |
|---|---|---|---|
| Autoregulation | EGR1 transactivates its own promoter via cis-element (−211/−203) | IL-1β-treated smooth muscle cells | Wang et al., 2010 |
| Histone modification | H3 phosphorylation and acetylation at Egr1 promoter | Chromatin regulation | Wang et al., 2010 |
Egr1, early growth response 1.
| Inducer | Mechanism | Effect | References |
|---|---|---|---|
| PMA | PKC activation → MAPK cascade | Rapid induction | Cheng et al., 1994 |
| Okadaic acid | PP1/PP2A inhibition | mRNA stabilization (t½ ≈ 2 h) | Cao et al., 1992 |
| Calyculin A | PP1/PP2A inhibition | Sustained post-transcriptional effect | Cao et al., 1992 |
| Serum | Growth factor stimulation | Short mRNA half-life (~12 min) | Cao et al., 1992 |
Egr1, early growth response 1; PMA, phorbol 12-myristate 13-acetate; MAPK, mitogen-activated protein kinase.
The abnormal or disease conditions also cause of Egr1 expression. These are including endoplasmic reticulum (ER) stress, cancer, osteoarthritis, cardiovascular conditions, neurodegenerative disorders, inflammatory diseases. For example, ER stress induce the transcription of Egr1 gene via the SRC-RAS-RAF-MEK-ERK signaling pathway, which enhances the phosphorylation of SRF and ELK1 and the binding to Egr1 promoter (Shan et al., 2019). In cancer the expression levels of EGR1 are high or low depending on the cancers. Such as RTK/MAPK/ERK/ELK1, PI3K/AKT, MAPKs (JNK/P38), FAK/RhoA/YAP-TAZ are known as an upstream signaling pathway activating Egr1 in tumorigenesis (Gitenay & Baron, 2009). In osteoarthritic cartilage a few cellular pathways are known to be involved (for example; FAK/Src/MAPK/ELK1/SRF, MAPK/AP-1/ELK1, beta-catenin). In cardiovascular disease, some signaling pathways such as FAK/Src/MAPK(ERK)/ELK1/SRF, MAPK/AP-1/ELK1, PKC/ROS/MAPK, or ELK/SRF are known (Khachigian, 2023). In the case of neurological diseases and inflammatory responses such as Alzheimer’s disease, chronic migraine, etc., a few signal transductions including Ca2+/MAPK(ERK1/2, p38)/ELK1/SRF, MAPK(ERK, JNK, p38)/AP1/ELK1, P38/JNK/AP-1, Ca2+/CaMKII/ERK/CREB/ELK1, MyD88/MAKP, are known for upstream signaling pathways for Egr1 expression (Bouallegue et al., 2013; Khachigian, 2023).
In addition, EGR-1 can regulate its own transcription. Histone H3 modification through phosphorylation and acetylation in the promoter of Egr1 gene are the key mechanisms in its expression regulation. EGR1 transactivates its promoter in smooth muscle cells exposed to interleukin 1beta (IL-1β) through a cis-acting element (–211/–203) (Wang et al., 2010).
Based on the signal transduction, Egr1 expression also induced artificially by the signal transfer mimicry likes phorbol 12-myristate 13-acetate (PMA) (Cheng et al., 1994). It also can be induced rapidly and transiently human and mouse fibroblast by the specific inhibitors of protein serine/threonine phosphatases 1 and 2A, okadaic acid and calyculin A, but sustained post-transcriptionally. Okadaic acid-induced Egr1 mRNA is significantly more stable (half-life: 2 h) than serum-induced Egr1 mRNA (half-life: 12 min) (Cao et al., 1992).
Translation of Egr1 mRNA is under the various levels in the known regulation mechanisms. In here we just introduce a few mechanisms among them. The metabolic factors can affect the translation of Egr1 mRNA. For example, insulin or glucose can induce the translation of Egr-1 protein (Hasan et al., 2003). Egr1 mRNA translation is sensitive to mTOR-elF4E related with 4E-BPs and elF4G1 as the effectors (Thoreen et al., 2012; Latancia et al., 2025). MAPK-MNK-eIF4E pathway enhances translation of Egr1 mRNA through regulation of inducible and constitutive elF4E phosphorylation (Waskiewicz et al., 1997; Ueda et al., 2004). miRNA such as miR-183, miR146a, miR-150 are involved in translational repression (Sarver et al., 2010; Contreras et al., 2015).
Egr1 protein is 533 amino acids with 57.5 kDa in human and 543 amino acids with 56.6 kDa in mouse. One of the most distinct structural features of EGR1 is the presence of three tandem C2H2-type zinc finger domains (Christy & Nathans, 1989; Hashimoto et al., 2014) (Fig. 5). These domains enable EGR1 to bind to specific GC-rich DNA motifs, particularly the consensus sequence 5’-GCG(T/G)GGGCG-3’ in the promoter region of target genes, regardless of the all different methylation states of CpGs (Hashimoto et al., 2014; Zandarashvili et al., 2015). Thereby, EGR1 regulates the expression of a wide array of target genes (Christy & Nathans, 1989; Hashimoto et al., 2014).
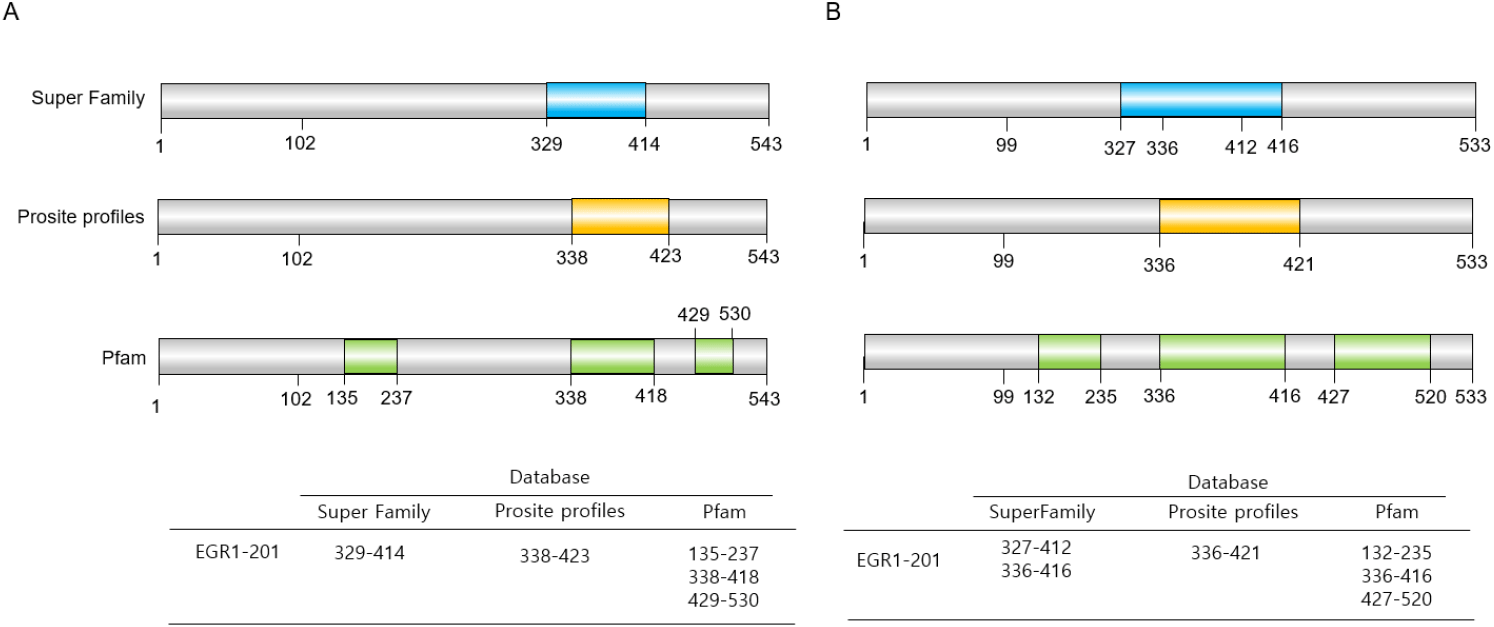
The activation of Egr-1 is disturbed over an extensive serine/threonine-rich N-terminal domain. Amino acids 281 to 314 repress the transcription on a heterologous DNA-binding domain. Specific DNA binding activity resides in the three zinc finger domains at its C-terminal region. Nuclear localization of Egr-1 is specified by signals in the DNA-binding domain and basic flanking sequences, a bipartite nuclear localization domain. In addition, a strong activating domain, a wake activation domain, and an inhibitory domain are known (Gashler et al., 1993) (Figs. 5 and 6). These zinc fingers have some similarity to the binding sites (5’-GGGGGGCGGGG-3’) of the Sp1 transcription factor. They share competing binding sites, G+C-rich elements in some genes, including platelet-derived growth factor (PDGF) A and B chains and adenosine deaminase (Khachigian et al., 1995).
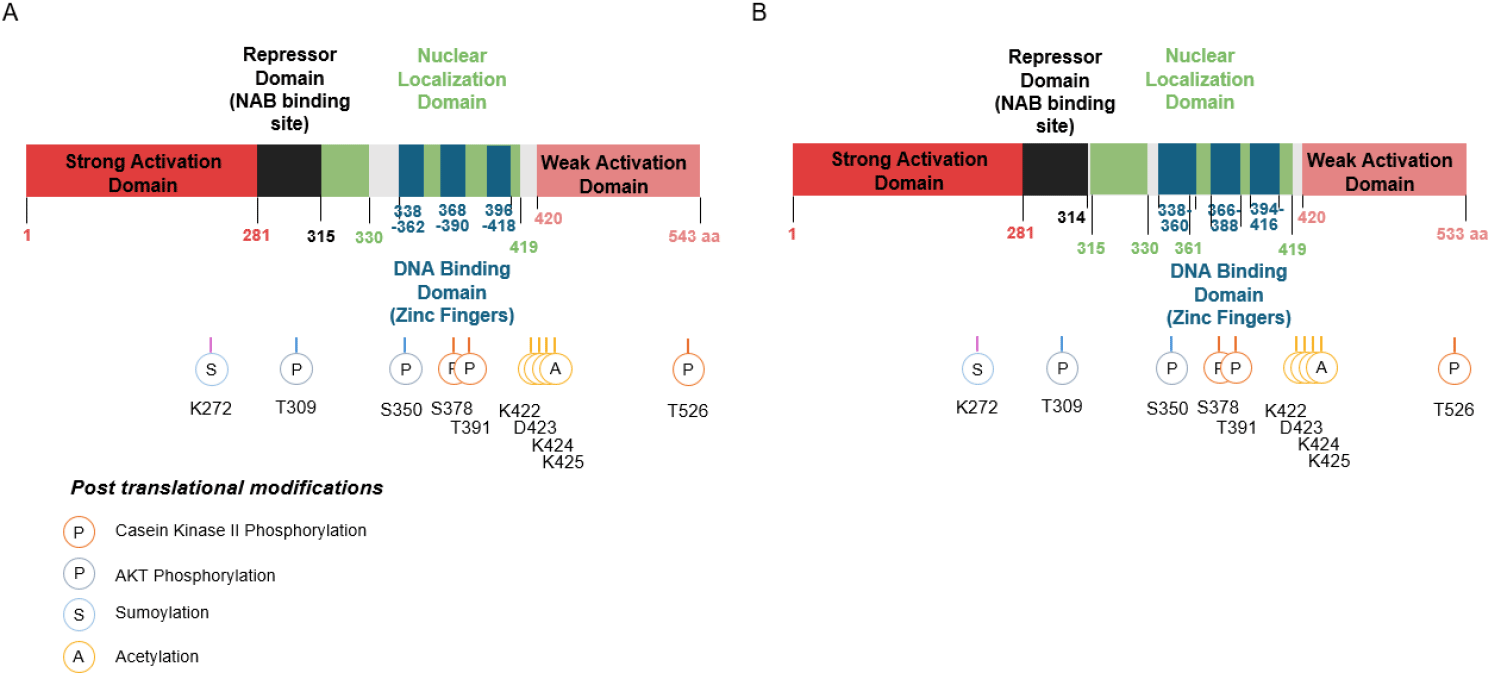
EGR1 interacts with NF-κB, AP-1, and SP1, forming cooperative or competitive transcriptional complexes in inflammatory signaling. These interactions determine whether EGR1 acts as a transcriptional activator or repressor, depending on chromatin state and epigenetic context (Trizzino et al., 2021). Moreover, chromatin remodeling plays an important role in EGR1-mediated transcription. EGR1 recruits cofactors such as CBP/p300 (acetyltransferases) or NuRD complex (repressors), which can either promote or silence transcription at specific loci, depending on the epigenetic marks present.
In addition to the structural factors that regulate activity, posttranslational modifications are also important in EGR1 (Fig. 6). Detection by Western blot analysis shows a range of 80 to 100 kDa, presumably due to post-translational modification of EGR1 (Cao et al., 1992; Gashler et al., 1993; Yu et al., 2009). The EGR1 domains are the target of phosphorylation by protein kinases and phosphatase (Cao et al., 1992). It is known that phosphorylation of EGR can either enhance or block EGR1 transcriptional activity. The Egr1 protein is weakly or not phosphorylated in quiescent cells, but multiple species of the phosphorylated forms of the Egr1 protein are detected in cells treated with either of the okadaic acid or calyculin A. Phosphorylation by the protein kinase Casein kinase II (CKII) in NIH3T3 cell suppresses transcription activity through inhibition of EGR1 DNA binding (Jain et al., 1996). Phosphorylation mediated by PKC and tyrosine kinase in response to UV damage in normal or immortalized cells has a role in a protective and anti-apoptotic function (Huang et al., 1998a), but Phosphorylation after unacetylation by UV-C irradiation in M12 cells leads to cell death by regulating target genes (Yu et al., 2004). EGR1 can be phosphorylated at S350 and T309 by Akt (Fig. 6), which promotes the interaction of EGR1 with the repressor domain of alternate reading frame (ARF) (Yu et al., 2009). EGR1 is sumoylated through the active p14ARF, which is under the Akt-EGR1-ARF-PTEN axis. EGR1 sumoylation is decreased by ARF reduction (Yu et al., 2009).
In addition to modification by phosphorylation, EGR1 is modified in other ways as follows. EGR1 can be acetylated in serum-stimulated prostate cells via CBP/p300 complexes, and this modification transactivates the survival genes, including itself. Acetylation stabilizes EGR1 and represses the negative feedback loop for Egr1 and p300/CBP (Yu et al., 2004). Acetylation of EGR1 is induced strongly by mastermind-like 1 (MAML1) through p300 in embryonic kidney (Hansson et al., 2012). EGR1 can be modified by ubiquitination. Ubiquitination inhibits the transcriptional activity of Egr-1 by the proteasome pathway. EGR1 is multi-ubiquitinated through its interaction with the proteasome component C8 (RC8) (Bae et al., 2002). Ubiquitination and sumoylation of EGR1 also induced by the small ubiquitin like modifier 1 (SUMO-1) and ubiquitin conjugating enzyme 9 (UBC9). It involved in stability of EGR1 in the human endothelial cell line ECV304 (Manente et al., 2011). Besides, EGR1 can be oxidized or bound with metals. The oxidized, or metal-free EGR1 does not bind to DNA (Huang & Adamson, 1993; Razmiafshari et al., 2001).
The effects of EGR1 vary depending on the cell type and context. Due to this property, as the initial wave of gene expression, EGR1 acts as a transcriptional “first responder,” linking external cues to downstream gene regulatory programs. Interestingly, the native antagonists are work in physiologically. N-terminal side of Zinc-finger DNA binding domain (repressor domain) interacts with NAB1 and NAB2 and these regulate negatively the EGR1 activity (Svaren et al., 1996; Swirnoff et al., 1998). These native antagonists are produced in the brain and other tissues as late genes and can regulate the EGR1 activity (O’Donovan et al., 1999; Hill et al., 2024; Yang et al., 2025). In addition, this gene is expressed by almost all the inducer. Therefore, it has been expected to be involved in development and physiology (Table 6), and this has been proven by numerous studies.
| Variables | Functional role | Key molecular targets / mechanisms | Phenotype (KO / OE) | References |
|---|---|---|---|---|
| Nervous system | Neurogenesis, synaptic plasticity, memory consolidation | Chromatin remodeling; hippocampal gene activation | KO: impaired long-term memory |
Beckmann & Wilce, 1997; O’Donovan et al., 1999; Rocks et al., 2023 |
| Connective tissue (tendon) | ECM regulation, tenogenic differentiation, repair | Col1a1, Col3a1, Col5a1, Scx, Tnmd; TGF-β2 signaling | KO: impaired ECM regulation; ↓ tendon strength |
Topilko et al., 1998; Lejard et al., 2011; Guerquin et al., 2013 |
| Bone / cartilage | Osteoclast regulation, fracture repair | CSF-1 suppression; β-catenin involvement; ECM remodeling | KO: ↑ bone resorption; fracture defects |
Srivastava et al., 1998; Reumann et al., 2011b |
| Adipose tissue | White vs beige adipocyte regulation | Lep, Ucp1 promoter binding; ECM gene repression | KO: spontaneous browning; protection from obesity |
Zhang et al., 2013; Milet et al., 2017 |
| Immune system (monocyte / macrophage) | Myeloid differentiation; inflammatory modulation | CSF1R regulation; NuRD recruitment; M1 polarization | Context-dependent inflammatory modulation |
Nguyen et al., 1993; Trizzino et al., 2021 |
| T cells | Differentiation, activation, exhaustion control | IL-2, IL-4, CD69 promoter binding; EGR1-NAB axis | KO: impaired differentiation; ↑ exhaustion |
Lohoff et al., 2010; Yang et al., 2025 |
| Female reproductive system | Decidualization, implantation, PR signaling | PR activation; Prl, Igfbp1 transcription | KO: infertility; impaired decidualization |
Kommagani et al., 2016; Kim et al., 2018 |
| Endocrine system | LHβ transcription; neuroendocrine regulation | Direct LHβ promoter binding; PC2 promoter sites | KO: infertility; LHβ deficiency | Jansen et al., 1997 |
| Fibroblasts / wound healing | Vascular remodeling | TGF-β modulation; ECM gene control | KO: reduced fibrosis; renal protection |
Ho et al., 2016; Li et al., 2021 |
| Angiogenesis | Vascular remodeling | VEGFR-1, VEGF, TGFB1, PDGF, PAI-1 regulation | Pathological angiogenesis in tumors |
Hasan et al., 2003; Li et al., 2019 |
| Cancer (context-dependent) | Tumor suppressor or promoter | TP53, PTEN activation; ferroptosis (Nrf2-HMOX1); PFKL suppression | Cancer-type specific outcomes |
Pagel & Deindl 2011; Lin et al., 2024; Pan et al., 2024 |
| Muscle | Myogenic differentiation | p57 (KIP2) regulation | Supports differentiation | Figliola et al., 2008 |
EGR1, early growth response 1; KO, knock-out; OE, over-expression; PR, progesterone receptor; PFKL, phosphofructoki-nase-1; PDGF, platelet-derived growth factor.
Egr1 expressed by the stimulation of growth factors in most cells leads to the activation of downstream pathways for growth. Besides, EGR1 can also regulate the amplitude of the expression rhythms of clock genes such as BMAL1, PER2, and NR1D1. As expected from such roles, Egr1 is involved in the development, homeostasis, and healing processes of tissues, including connective and nervous tissues (O’Donovan et al., 1999; Havis & Duprez, 2020). EGR1 regulates responses to various stimuli, as previously mentioned. It plays a crucial role in regulating cell survival, proliferation, and cell death. Those are including control of neural cell death (Xie et al., 2011), neuronal plasticity (Wei et al., 2000; Lee et al., 2004), proliferation (Mayer et al., 2009), T cell differentiation (Safford et al., 2005), monopoiesis (Nguyen et al., 1993), myelination (Topilko et al., 1994), inflammation (Ji et al., 2003), ossification (Levi et al., 1996), muscle spindle formation (Tourtellotte & Milbrandt, 1998), and synthesis such as LHβ or phenylethanolaine-N-methyltransferase (Lee et al., 1996; Morita et al., 1996; Wolfe & Call, 1999). It encompasses a broad spectrum of developmental processes across multiple systems, including the brain, reproductive organs, immune tissues, and cardiovascular system. Its expression begins early in embryogenesis, contributing to lineage differentiation and tissue patterning during development.
EGR1 orchestrates the changes in gene expression that underlie neural plasticity during neurogenesis, including synaptic plasticity (O’Donovan et al., 1999; Rocks et al., 2023). In the nervous system, EGR1 contributes to synaptic plasticity and memory consolidation, particularly in the hippocampus. Egr1-deficient mice exhibit impaired long-term memory and learning, indicating its critical role in cognition (Beckmann & Wilce, 1997). Overexpression of EGR1 induces sex-specific changes in ventral hippocampal neuronal synaptic plasticity by opening neural chromatin with hippocampus-dependent behaviors (Rocks et al., 2023).
In connective tissue genesis, the extracellular matrix is a main component of connective tissue. During connective tissue genesis, Egr1 primarily functions through the regulation of the extracellular matrix (Havis & Duprez, 2020). Egr1 null mouse embryo fibroblasts bypass replicative senescence and exhibit a loss of DNA damage response and an apparent immortal growth through the loss of p53 function. In Egr1 null embryo fibroblasts, 266 transcripts are differentially expressed more than 2-fold, and one of the direct target genes is p53 (Krones-Herzig et al., 2005).
In Egr1 knockout mice models, various defects in connective tissue are evaluated. Egr1 null mutations affect the formation and homeostasis of tendon, cartilage, bone, and adipose tissue (Topilko et al., 1998). In vertebrate tendon formation, the involvement of EGR1 is known. Its expression is correlated with increased collagen production during tendon cell differentiation. Egr1 knockout mice do not display a strong overt tendon phenotype with the loss of tendon-associated collagens (Col1a1, Col3a1, Col5a1, Col12a1, and Col14a1) expression (Lejard et al., 2011). Egr1 downregulation is associated with a loss of the tenogenic differentiation potential in ageing human tendon progenitor cells, while EGR1 gain-of-function has the ability to rescue tendon differentiation potential (Han et al., 2017). In tendon stem cells, EGR1 induces tenogenic differences (Tao et al., 2015). In addition, Egr1 is required for the correct expression of matrix genes such as Col1a1, Col1a2, Scx, Tnmd, Col5a1TTnc, and Dcn, during tendon differentiation and repair. The application of EGR1-producing MSCs increases the formation of tendon-like tissues in Achilles tendon injury, partially mediated by TGF-β2, and increases the strength of the healing tendon (Guerquin et al., 2013; Hammerman et al., 2014; Havis & Duprez, 2020).
The Egr1 gene is expressed in the endochondral processing area and periosteal region of the long bone during embryonic development through Krüppel-like factor 5 and β-catenin (McMahon et al., 1990; Sun et al., 2019). In EGR1 knockout mice, bone resorption is increased through the positive regulation of osteoclast differentiation (Srivastava et al., 1998). EGR1 suppresses the osteoclastogenic cytokine CSF-1 production by stromal cells (Srivastava et al., 1998). On the other hand, Egr1 deficiency leads to persistent accumulation of fibrin in the endochondral bone fracture repair, abnormal sectional geometry, and defects in extracellular matrix regulation for load and stiffness (Reumann et al., 2011a,b).
It is involved in the induction of myeloid cell differentiation along the monocyte lineage and in the activation of monocytes (Kharbanda et al., 1991; Nguyen et al., 1993; Pham et al., 2012). EGR1 regulates monocyte developmental genes such as CSF1R (Trizzino et al., 2021). In macrophages, EGR1 suppresses inflammatory enhancer activity by recruiting the NuRD corepressor complex, acting as a brake on chronic inflammation (Trizzino et al., 2021). EGR1 is involved in T cell differentiation and T cell activation. The selective ablation of Egr1 in CD4+ T cells impedes the differentiation (Yang et al., 2025). Upregulating EGR1 activity and expression reduces CAR-T cell exhaustion and blocks exhausted T cell terminal differentiation (Sui et al., 2024). Egr1 expression upon T cell stimulation is occurs predominantly in T helper type 2 (Th2) compared with type 1 (Th1) cells. EGR1 binds to the IL-4 promoter upon T cell stimulation and is involved in the acute phase of L-4 transcription (Lohoff et al., 2010).
In fat cell differentiation, EGR1 is also an important task force (TF). Egr1 is directly recruited to the leptin gene (Lep) and uncoupling protein 1 (Ucp1) gene, markers of white adipocyte and beige adipocyte markers, respectively (Milet et al., 2017; Mohtar et al., 2019). Egr1 is induced by insulin in adipocytes and results in the increase of leptin transcription by directly binding to the promoter of leptin (Mohtar et al., 2019). Egr1 deletion causes an increase in spontaneous browning of subcutaneous white adipose tissue along with downregulation of ECM genes such as Col1a subfamilies, Fn1, Dcn, and Periostin (in hypodermal adipose tissue (Milet et al., 2017). Egr1−/− mice are protected from diet-induce obesity and obesity-associated pathologies such as fatty liver, insulin resistance and hyperlipidemia (Zhang et al., 2013). Egr1 overexpression in C3H10T1/2 cells prevents the beige adipocyte differentiation (Milet et al., 2017).
In the female reproductive system, EGR1 plays a pivotal role in endometrial decidualization and embryo implantation. In uterine endometrium, EGR1 is induced by estrogen and it promotes the expression of progesterone receptor (PR). Egr1 knockout causes defective PR signaling in epithelium and poor communication with stromal cells (Kim et al., 2018). It also suggested that EGR1 directly promotes the transcription of Prl and Igfbp1, and its deficiency in endometrial stromal cells results in impaired decidual response (Kommagani et al., 2016). Additionally, EGR1 has been implicated in age-associated follicular atresia. In aging ovaries, Egr1 expression increases and contributes to granulosa cell apoptosis via NF-κB signaling (Yuan et al., 2016), suggesting a role in ovarian reserve decline.
EGR1 also contributes to angiogenesis and vascular remodeling by regulating genes like vascular endothelial growth factor receptor-1(fit-1), vascular endothelial growth factor, transforming growth factor beta1, PDGF, plasminogen activator inhibitor-1 (Hasan et al., 2003). Interestingly, the angiogenic role of EGR1 can observed in pathological conditions through upregulation of pro-tumorigenic factors (Li et al., 2019). In addition to these, it is also suggested that EGR1 is involved in muscle cell differentiation through the regulation of p57 (KIP2) expression (Figliola et al., 2008).
The Egr1 gene is translated in various tissues, including epithelium (epithelial cells), connective tissues (tendon, cartilage and bone, and adipose tissue) (Havis & Duprez, 2020), nervous system, such as forebrain and neurons (Milbrandt, 1987), fibroblast, and lymphocyte (Sukhatme et al., 1988). However, its expression mechanism depends on the general stimulation, it is better to say it can be expressed all the cells. So it is believed that EGR1 has multiple physiological functions.
As mentioned, EGR1 functions as a context-dependent transcription factor with a broad range of physiological roles depending on the stimuli. One of the roles of EGR1 is the function in the endocrine system. Since Egr1 knockout mice are infertile due to defects in endocrine, it is suggested that EGR1 is involved in regulating neuropeptide gene expression (Jansen et al., 1997; O’Donovan et al., 1999; Xu et al., 2022). Egr1 knockout mice are deficient in LHβ. LHβ gene is downstream of EGR1 and it has EGR response element. Prohormone convertase 2 (PC2) promoter shows direct neuroendocrine-specific expression of the luciferase reporter gene and PC2 promoter has two EBS (Jansen et al., 1997).
In fibroblasts, EGR1 regulates wound healing-related genes (Yeo et al., 2020). Egr1−/− mice display reduced TGF-beta activity and reduced renal fibrotic zones and are protected from renal failure (Ho et al., 2016). It is also well known that the functions of EGR1 in ECM-related pathological conditions. In fibrotic conditions, Egr1 is associated with the progression of the abnormal production of ECM (Havis & Duprez, 2020). In the renal fibrosis of diabetic kidney disease, KLOTHO, an antiaging protein, attenuates renal fibrosis along with preventing epithelial-to-mesenchymal transition in part by Egr1 downregulation (Li et al., 2021).
In cancer, it is known that EGR1 has been associated with carcinogenesis. EGR1 can act as a tumor suppressor (some of them are via TP53, PTEN) or a tumor promoter depending on cancer type (Pagel & Deindl, 2011; Magee & Zhang, 2017). EGR1 activates expression of p53/TP53 and TGFB1, and thereby helping prevent tumor formation (Virolle et al., 2003; Li et al., 2019). In colon cancer cells, EGR1 overexpression inhibits cell proliferation, migration, and invasion. The knockdown of Egr1 increases colon cancer cell proliferation, migration, and invasion, along with the expression of cyclin-dependent kinase-like 1 (Shao et al., 2021). On the other hand, Egr1 also suppresses the growth or proliferation of various cancers or cancer cell lines. In human fibrosarcoma cell, HT1080, EGR1 transfection suppresses the proliferation through Bcl2 expression (Huang et al., 1998b). In hepatocellular carcinoma, EGR1 suppresses cell proliferation through phosphofructokinase-1 (PFKL, liver type) downregulation (Pan et al., 2024). In breast cancer cells, Egr1 promotes erastin-induced ferroptosis through activating the Nrf2-HMOX1 signaling pathway (Lin et al., 2024).
In immune function, EGR1 is a context-dependent modulator. It can promote early inflammatory gene waves before Nf-kB-dependent sustained programs dominant in macrophages and monocytes (Trizzino et al., 2021; Zhang et al., 2024). It induces macrophage polarization toward an M1 phenotype (Wang et al., 2023). EGR1 suppresses the phagocytosis of P. aeruginosa by macrophages. EGR1 upregulates autophagy and inhibits NRF2 signaling (Pang et al., 2022). The neutrophils expresse EGR1 as a result of responding to cytokines, and it is implicated in vascular inflammation and leukocyte recruitment (Schmidt et al., 2008; Cullen et al., 2010; Grieshaber-Bouyer et al., 2021). In dendritic cells, EGR1 links innate sensing to adaptive priming (ten Hoeve et al., 2019). In T cells, EGR1 is involved in early activation through binding to the promoters of IL2, CD69 and AP-1 cooperative targets. It influences on activation-induced cell fate decisions and EGR1-NAB regulator axis (Decker et al., 1998; Kumbrink et al., 2005; Collins et al., 2008). In autoimmune and inflammatory disorders, EGR1 contributes to chronic cytokine expression, pathological macrophage activation, T cell differentiation, and vascular inflammation (Yang et al., 2025). Egr1 expression acts as a cytokine storm initiation phase (Liu et al., 2016; Zhao et al., 2021).
As a transcription factor, EGR1 regulates a broad range of target genes depending on cell type, stimulus, and developmental stage. Its target genes are involved in diverse biological pathways as mentioned above. In addition to the previously mentioned targets, many target genes are evaluated.
The well-known downstream genes include p53/TP53 and TGFB1 (helps prevent tumor formation), IL1B and CXCL2 (inflammatory processes and development of tissue damage after ischemia), LBH (hormonal secretion), BMAL1, PER2, and NR1D1 (regulate the amplitude of the expression rhythms of clock genes). EGR1 directly binds to the TP53 promoter and induces its transcription, thereby contributing to cell cycle arrest and apoptosis in stress conditions (Thiel & Cibelli, 2002). Similarly, PTEN, a key negative regulator of the PI3K/AKT pathway, is also transcriptionally upregulated by EGR1 in cancer and metabolic tissues mediated by Akt-EGR1-ARF-PTEN axis (Yu et al., 2009).
VEGF, FGF2, and TGF-β1, which control endothelial cell migration and vessel formation, are known targets of EGR1. These functions are especially relevant in wound healing, tissue repair, and cancer progression (Li et al., 2019). Additionally, EGR1 also participates in feedback regulatory loops of transcription factors. For example, EGR1 can activate Egr2 and Egr3, other early response genes in the same family, as well as transcription factors such as JUN, FOS, and NAB2, establishing a broader transcriptional network. In some systems, EGR1 is also known to induce its own expression transiently, forming a short-lived auto-regulatory loop.
In addition, extracellular matrix components (Acan, Bgn, Cols, Dcn, Fbn1, Fn1, etc.), secreted proteins and hormones (Bglap, Csf10, Ctsk, Lep, Tgfb2, Tnmd, etc.), cytoplasmic proteins (Pnpla2, Ucp1, etc.), transcription factors/nuclear proteins (Cebpb, Foxc2, Mkx, Pparg, Scx, etc.) (Gaut et al., 2016; Havis & Duprez, 2020). In addition to them, CSF1R and IL-4 in inflammation, and apoAI and LDLRin metabolism are also suggested.
CONCLUSION
Egr1 is a rapidly induced immediate-early transcription factor whose biological function varies significantly depending on the environment, yet its core genetic structure is remarkably well conserved between humans and mice. In both species, the Egr1 gene consists of two exons and one intron, encoding a highly conserved C2H2-type zinc finger DNA-binding domain. This structural conservation supports the maintenance of core transcriptional functions, while species-specific differences in promoter usage and enhancer landscapes allow for diverse cis-regulation. These regulatory features, along with multiple modulations at the transcriptional, posttranscriptional, translational, and posttranslational levels, highlight the environmentally dependent behavior of Egr1, acting as either an activator or a repressor depending on the tissue.
As discussed above, Egr1 possesses a structurally simple and highly conserved gene, but its expression is meticulously controlled by various cis-regulatory elements and multilayered regulatory mechanisms. These genetic anatomy characteristics provide a molecular basis for Egr1 to function as a key regulatory factor across multiple organs and systems, including development, homeostasis, and tissue healing, rather than being restricted to specific tissues or a single physiological system.
