INTRODUCTION
The vertebrate kidney is an essential excretory organ that maintains body fluid and electrolyte balance, removes metabolic and xenobiotic wastes, and produces hormones vital for systemic homeostasis. During embryogenesis, paired kidneys arise from the intermediate mesoderm and develop through three successive stages, the pronephros, mesonephros, and metanephros, exhibiting increasing morphological and functional complexity (Wessely & Tran 2011). In amphibian embryos, the pronephric kidney functions as the initial excretory organ, operating independently during early development, and is later replaced by the mesonephros. Conversely, in mammals, the pronephros and mesonephros are transient precursors, whereas the metanephros gives rise to the definitive adult kidney (Cirio et al., 2011). Despite interspecies differences, all renal forms share a conserved structural and functional unit - the nephron-comprising the glomerulus, segmented tubule, and collecting duct (Chae et al., 2017; Corkins et al., 2023).
Xenopus laevis is a useful vertebrate model for studying nephrogenesis because its embryonic pronephros resembles key aspects of core mammalian nephron architecture and gene expression programs. Its external development enables direct imaging and precise microinjection, whereas rapid organogenesis, large clutch sizes, and a fully sequenced genome facilitate in vivo genetic and ‘omics’ approaches (Schmitt et al., 2014; Lienkamp 2016). Thus, the Xenopus pronephros offers a simplified yet highly conserved system for investigating the molecular mechanisms involved in kidney development and disease.
The proper execution of nephrogenesis, involving coordinated cell proliferation and differentiation, requires precise metabolic regulation and robust defense against oxidative stress (Chae et al., 2017). Isocitrate dehydrogenases (IDHs) play a central role in maintaining this balance. The IDH family comprises three vertebrate isoforms, Idh1, Idh2, and Idh3, which differ in subcellular localization and cofactor usage. Idh1 is a cytosolic/peroxisomal, NADP+-dependent enzyme; Idh2 is a mitochondrial, NADP+-dependent enzyme; and Idh3 is a mitochondrial, NAD+-dependent enzyme participating in the TCA cycle (Xu et al., 2004). In addition to their roles in central carbon metabolism, Idh1 and Idh2 supply α-ketoglutarate (α-KG) and NADPH to support redox balance and biosynthesis.
Pathologically, recurrent heterozygous mutations at conserved catalytic residues, R132 in Idh1 and R140/R172 in Idh2, confer a neomorphic activity that reduces α-KG to D-2-hydroxyglutarate (2-HG). This competitively inhibits α-KG–dependent dioxygenases, disrupting epigenetic regulation and cellular differentiation. Such mutations are frequent in diffuse gliomas, acute myeloid leukemia (AML), and cholangiocarcinoma (Dang et al., 2009; Ward et al., 2010; Losman & Kaelin 2013). However, the role of IDH isoforms in early vertebrate organogenesis, including kidney development, remains largely unexplored.
To address this gap in the literature, this study aimed to investigate the roles of Idh1 and Idh2 in pronephric development in X. laevis. We analyzed their temporal and spatial expression, performed morpholino-based loss-of-function analyses, and quantified pronephric defects using tubule markers. Furthermore, rescue assays showed that idh1*R132H restored pronephric development in idh1-depleted embryos, suggesting a context-dependent developmental function. These findings identify Idh1 and Idh2 as key regulators of pronephric morphogenesis and provide novel developmental insight into disease-associated Idh1 mutations.
METHODS
Mature X. laevis frogs were obtained from the Korean Xenopus Resource Center for Research and maintained in plastic aquaria with continuously circulated, dechlorinated water at 18°C, in accordance with the recommendations of the Institutional Review Board of Kyungpook National University (Korea). Ovulation in adult females was induced by injecting 1,000 IU of human chorionic gonadotropin (hCG; Sigma-Aldrich, St. Louis, MO, USA) into the dorsal lymph sac on the evening prior to the experiment. After approximately 16 h, eggs were collected in 1× MBS solution (88 mM NaCl, 5 mM HEPES, 2.5 mM NaHCO3, 1 mM KCl, 1 mM MgSO4, and 0.7 mM CaCl2; pH 7.8) by gently squeezing the females. Eggs were fertilized in vitro using a sperm suspension prepared from freshly dissected testes. After several rinses with 0.1× MBS, the eggs were mixed with the sperm solution to allow fertilization. The jelly coats were removed by swirling the embryos in 2% L-cysteine (pH 7.8) prepared in 0.5× MBS. Following dejellying, embryos were washed several times with 0.5× MBS and cultured at 15°C–18°C until they reached the desired developmental stages.
The coding sequences of idh1.L and idh2.L (NCBI accession nos. NM_001094553.1 and XM_018253189.2, respectively) were amplified through polymerase chain reaction (PCR) using X. laevis cDNA. HA-tagged idh1.L and idh2.L fragments were subcloned into the pCS107 expression vector and linearized with Apa I (Takara Bio, Shiga, Japan) for in vitro transcription. Capped mRNAs were synthesized using the SP6 mMESSAGE mMACHINE Kit (Invitrogen, Carlsbad, CA, USA). To generate morpholino-resistant constructs for rescue assays, idh1 mRNAs (designated idh1*) containing six silent point mutations near the start codon were produced by site-directed mutagenesis. Additionally, a site-specific mutant (idh1*R132H) was generated using the same approach. Translation-blocking antisense morpholino oligonucleotides (MOs) targeting the start codon regions of idh1.L (5'-TAATTTTCTTGGACATCCTGGCGTT-3') and idh2.L (5'-TAACAGCACGCAGATAACCTGCCAT-3') were designed and synthesized by Gene Tools LLC (Philomath, Oregon, OR, USA). Synthesized mRNAs or MOs were microinjected into one-cell stage embryos, which were subsequently cultured in 0.5× MBS at 23°C until they reached the desired developmental stages.
Embryos were fixed at the desired developmental stages in MEMFA solution (4% paraformal-dehyde, 0.1 M MOPS [pH 7.4], 1 mM MgSO4, and 2 mM EGTA) overnight at 4°C. DNA templates for RNA probe synthesis were linearized using the appropriate restriction enzymes, and DIG-labeled antisense RNA probes were transcribed in vitro using SP6 or T7 RNA polymerase (Invitrogen). Hybridized probes were detected using an alkaline phosphatase-conjugated anti-digoxigenin antibody (1:1,000; Roche, Mannheim, Germany) and visualized using an NBT/BCIP substrate solution (Roche), following previously described procedures (Kim et al., 2025). Whole-mount in situ hybridization (WISH) was performed following standard protocols using probes specific for idh1.L (NM_001094553.1), idh2.L (XM_018253189.2), smp30.L (NM_001085655.1), and atp1a1.L (NM_001090595.1).
Total RNA was isolated from X. laevis embryos using an RNA Lysis Reagent (5 Prime, Hilden, Germany). Complementary DNA (cDNA) was synthesized from 1 μg of total RNA using the PrimeScript™ RT Reagent Kit (Takara), according to the manufacturer’s instructions. Reverse transcription–polymerase chain reaction (RT-PCR) was performed with gene-specific primers, and amplified products were separated on 1% agarose gels. Bands were visualized using a gel imaging system (WiseCapture I-1000; Daihan Scientific, Seoul, Korea). Expression levels were normalized against odc, which served as the internal control.The sequences of all primers used for RT-PCR are listed in Table 1.
RT-PCR, reverse transcription–polymerase chain reaction.
All experiments were conducted with at least three independent biological replicates. Quantification of embryo dimensions and WISH signal intensity was performed using ZEN 3.7 software (Carl Zeiss, Oberkochen, Germany). Statistical analyses were performed using GraphPad Prism version 9.0 (GraphPad Software, San Diego, CA, USA). Data are expressed as the mean ± SEM. Statistical significance between groups was assessed using one-way analysis of variance (ANOVA) followed by Dunnett’s multiple comparisons test. Significance levels were defined as *p<0.05, **p<0.01, ***p<0.001, and ****p<0.0001.
RESULTS AND DISCUSSION
To investigate the potential roles of Idh1 and Idh2 during X. laevis embryogenesis, their temporal expression profiles were analyzed via RT-PCR using staged embryos. Both idh1 and idh2 are maternally expressed genes; however, their developmental expression dynamics differed significantly. idh1 expression increased at the blastula stage (stage 8), decreased during gastrulation (stage 12), and remained low through stage 20. From stage 25 onward, expression gradually increased, reaching higher levels at stages 30 and 40 (Fig. 1A and B). In contrast, idh2 expression remained relatively constant from stage 8 through stage 40, indicating a stable expression pattern throughout development (Fig. 1A and B).
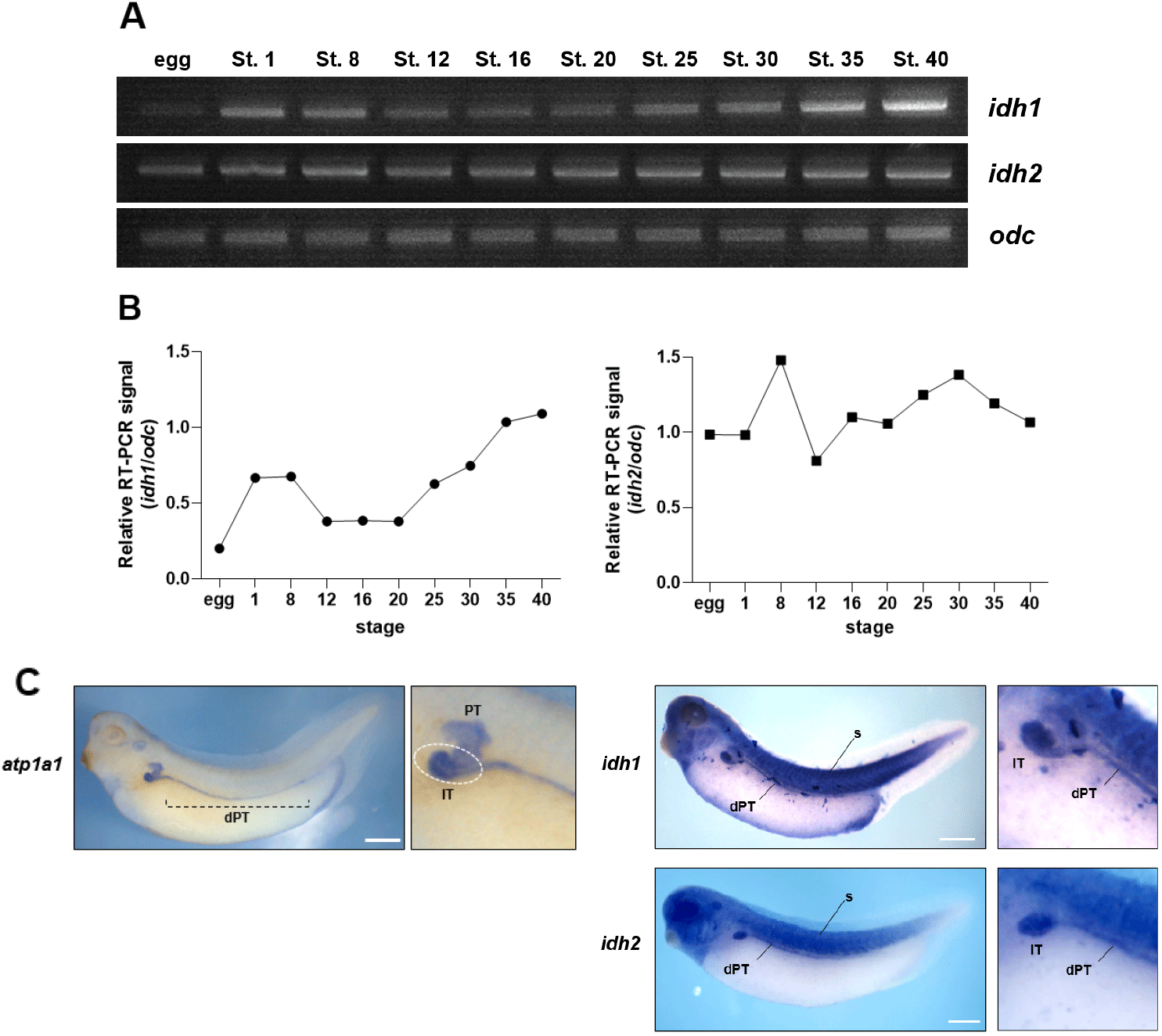
To determine spatial distribution, WISH was performed using late tailbud-stage embryos. Both genes exhibited partially overlapping expression domains, with strong signals detected in the craniofacial region, somites, and the pronephric kidney (Fig. 1C). Within the pronephros, expression was predominantly observed in the intermediate tubule (IT) and extended posteriorly along the distal pronephric tubule (dPT), as determined by comparison with the pronephric marker atp1a1 (Fig. 1C). The specific localization of idh1 and idh2 in these metabolically active tissues suggests that both genes may play important roles in X. laevis embryonic development.
To further assess the developmental roles of Idh1 and Idh2, loss-of-function experiments were conducted by injecting antisense MOs (60 ng) into one-cell stage Xenopus embryos. Given that both genes are expressed in the pronephric region (Fig. 1B), we assessed kidney development via WISH using the proximal tubule marker smp30 and the pan-pronephric marker atp1a1. The results revealed that expression of both smp30 and atp1a1 was reduced in morphants, with more prominent effects observed in idh1 morphants than in idh2 morphants. Particularly, the proximal tubule appeared less developed in idh1 morphants compared with control MO-injected embryos (Fig. 2A). Quantitative analysis further showed that the proportion of embryos exhibiting defective expression patterns was significantly higher in idh1 morphants (Fig. 2B and C). Although early nephric transcription factors such as pax8 were not examined, the reduced expression and incomplete structural organization of pronephric markers suggest that idh1 and idh2 are more likely involved in pronephric morphogenesis than in early specification.
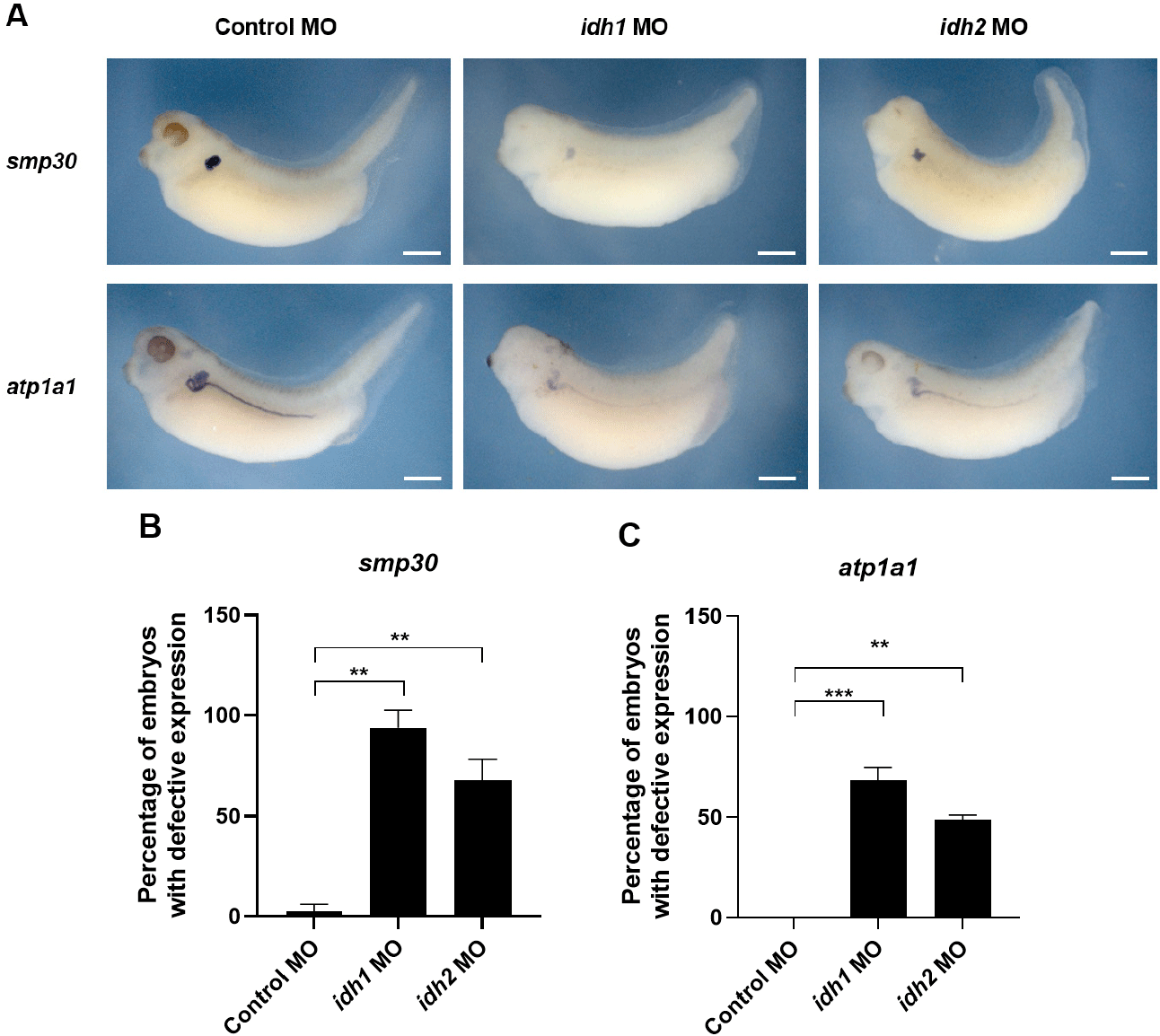
Previous studies have demonstrated that interactions between developing somites and the pronephros influence the patterning of both tissues (Vize et al., 1995; Mauch et al., 2000; Guillaume et al., 2009). Consistent with these findings, idh1 and idh2 were also expressed in somites, and their morphants exhibited dorsal curvature (Fig. 2A), suggesting that abnormal somite formation may secondarily contribute to the pronephric defects observed.
Collectively, these findings indicate that Idh1 and Idh2 are required for proper pronephric development during Xenopus embryogenesis.
Idh1 contains large and small domains that form an active-site cleft for binding isocitrate and NADP+ (Xu et al., 2004). The conserved arginine at position 132 (R132), located in a loop of the small domain, coordinates the β-carboxyl group of isocitrate during catalysis. Substitution of this residue with histidine (R132H) disrupts substrate binding and converts Idh1 into a neomorphic enzyme that reduces α-KG to D-2-hydroxyglutarate (2-HG), a metabolite influencing epigenetic reprogramming in human cancers (Dang et al., 2010; Ward et al., 2010; Krell et al., 2011).
To assess whether this oncogenic mutation influences Idh1 function during embryonic kidney formation, we tested whether idh1*R132H could rescue pronephric defects caused by idh1 knockdown in X. laevis embryos. Co-injection of wild-type (WT) idh1* mRNA efficiently restored the expression of the pronephric marker atp1a1, confirming the specificity of the MO-induced phenotype (Fig. 3A). Notably, idh1*R132H mRNA achieved a comparable level of rescue (Fig. 3A and B), suggesting that the R132 residue, although critical for catalytic activity in mammalian metabolism, may not be strictly required for Idh1’s developmental contribution to pronephric morphogenesis.
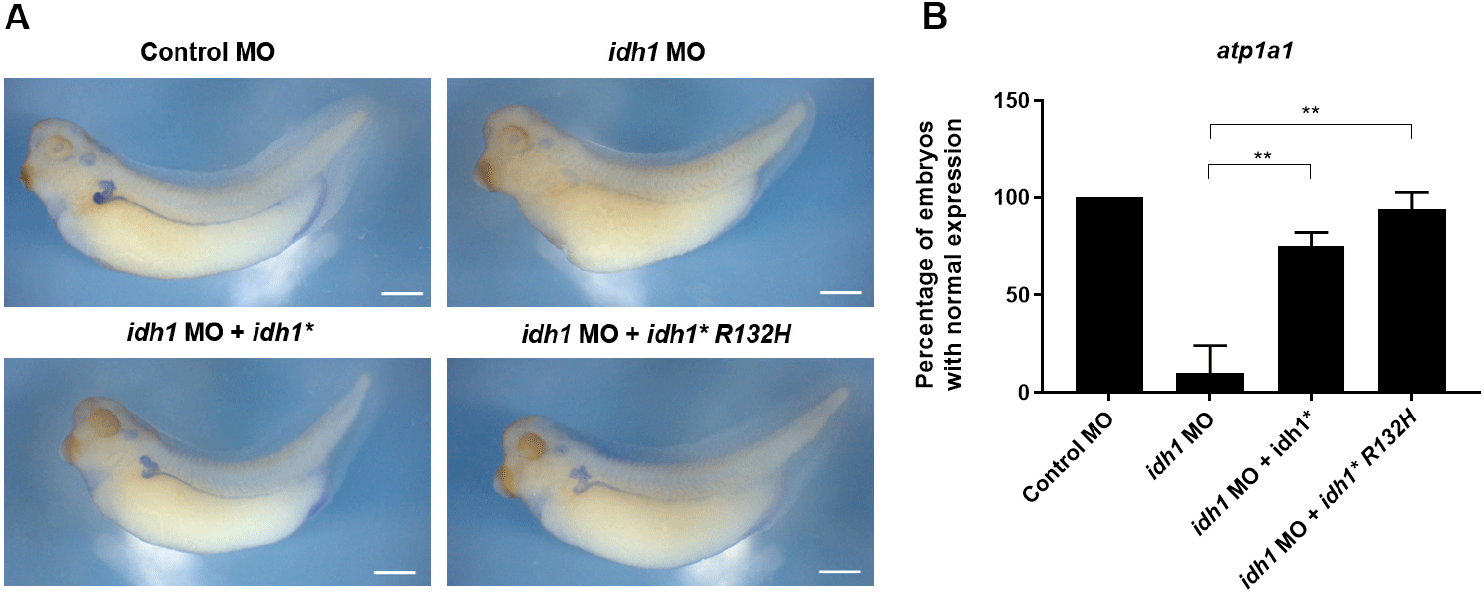
Alternatively, embryonic metabolic conditions may suppress the oncogenic activity of the R132H variant, thereby minimizing its effect on pronephric development. Consistent with reports that mutant Idh1 alone does not transform normal cells and acts in a context-dependent manner (Chaturvedi et al., 2013; Bardella et al., 2016), our findings suggest that its developmental effects may be shaped by tissue-specific metabolism. Measuring 2-HG levels in embryos overexpressing idh1*R132H would further clarify this possibility.
In conclusion, our findings demonstrate that Idh1 and Idh2 are necessary for proper pronephric development in X. laevis embryos, and that the oncogenic idh1*R132H variant can restore pronephric defects in idh1-depleted embryos. These findings highlight a context-dependent developmental role for Idh1 that is distinct from its canonical metabolic function.
