INTRODUCTION
In developmental biology applications, somatic cell nuclear transfer (SCNT) is an important process for producing cloned animals and specific pluripotent embryos or cells. It is widely used in the production of transgenic animals, xenotransplantation, and animal cloning, but one of the major deficiencies of the technique is the low efficiency of the reprogramming of the SCNT embryos (Onishi et al., 2000). The incomplete nuclear reprogramming of the SCNT embryos leads to epigenetic modification (Vignon et al., 2002), aberrant gene expression (Kang et al., 2001), and abnormal X-chromosome inactivation (Xue et al., 2002; Nolen et al., 2005) resulting in abnormal offspring (Kubota et al., 2000; Vignon et al., 2002).
Several studies have sought to improve the nuclear reprogramming by treating with a DNA methylation agent (Enright et al., 2003), impeding Xist expression from the active X-chromosome (Inoue et al., 2010), and modulating nuclear remodeling (Kwon et al., 2008). However, most of these experimental approaches were suitable only for the in vitro culture of SCNT embryos. The SCNT embryos suffered physical and chemical stresses by manipulation during the SCNT process, including micromanipulation, electrofusion, and activation, which restricted nuclear reprogramming (Hwang et al., 2013). It was reported that SCNT technique could alter the mitochondrial membrane potential and trigger the fluctuation of cytosolic Ca2+ concentration (Spikings et al., 2006). Our previous studies were about the physical and chemical stresses of the oocytes caused by the reactive oxygen species (ROS) released during SCNT (Hwang et al., 2013; Bae et al., 2015). The physical and chemical stresses during SCNT resulted in more ROS being released damaging the mitochondria and DNA (Hwang et al., 2013). Antioxidant treatment during SCNT inhibited the release of ROS thereby reducing cellular damage and enhancing the reprogramming of the SCNT embryos (Bae et al., 2015). Besides ROS, cytoplasmic endoplasmic reticulum (ER) stress can induce cellular damage including apoptosis (Tabas & Ron, 2001; Groenendyk & Michalak, 2005; Szegezdi et al., 2006) and mitochondrial dysfunction (Wu et al., 2015), which can affect embryo development (Zhang et al., 2012a; Yoon et al., 2014). The most important functions of the ER are protein synthesis, secretory modification and protein folding into the native conformation. Disturbed ER homeostasis affects proper protein folding and leads to the accumulation of unfolded or misfolded proteins in the ER lumen. When the amount of unfolded proteins exceeds the folding capacity of the ER, it results in ER stress. In this condition, cells activate a defense mechanism called the unfolded protein response (UPR) or ER stress response (Boyce & Yuan, 2006; Malhotra & Kaufman, 2007). Malfunction of the ER stress response caused by aging, genetic mutations, or environmental factors can result in several diseases like diabetes, inflammation, and neurodegenerative disorders (Yoshida, 2007).
Alleviation of ER stress reduced apoptosis and enhanced the development of bovine SCNT embryos (Song et al., 2014). Expression of the ER stress-associated gene and apoptotic gene was high in SCNT blastocysts (Cánepa et al., 2014). ER stress is closely related to the release of ROS (Malhotra et al., 2008; Yoon et al., 2014). As ROS were released during the SCNT process also, they induced ER stress. Therefore, ER stress should be inhibited at the initial step of the SCNT process rather than during the culture period to prevent the ER stress-derived cellular damage. Thus, in this study, we examined the ER stress and subsequent apoptosis in porcine SCNT embryos and compared them with in vitro fertilization (IVF) embryos.
MATERIALS AND METHODS
All chemicals and reagents were purchased from Sigma-Aldrich Chemical Co. (St Louis, Mo, USA) unless otherwise indicated.
Porcine ear skin fibroblast cells were seeded in a 24-well plate and cultured in Dulbecco’s modified Eagle’s medium (DMEM; Gibco, Grand Island, NY, USA) supplemented with 10% fetal bovine serum (FBS; GenDEPOT, Katy, Texas, USA) and 1% penicillin and streptomycin (P/S; Nalgene, Manassas, Virginia, USA) at 39℃, 5% CO2 in air for 3-4 days. To induce ER stress, cells that reached 70-80% confluence were treated with 2 μg/mL or 5 μg/mL tunicamycin (TM), an ER stress inducer, for 3 h or 6 h, respectively. The cells were assessed for ER stress after the treatments.
Porcine cumulus-oocyte complexes (COCs) were collected from the follicles (3-6 mm diameter) and cultured in 500 μL droplets of in vitro maturation (IVM) medium overlaid with paraffin oil at 39℃, 5% CO2 in air for 42-44 h. The IVM medium was Tissue Culture Medium 199 (TCM199; Gibco) supplemented with 0.1% polyvinyl alcohol (PVA), 3.05 mM D-glucose, 0.91 mM Na-pyruvate, 75 μg/mL penicillin G, 50 μg/mL streptomycin, 0.57 mM cysteine, 10 ng/mL epidermal growth factor, 0.5 μg/mL luteinizing hormone, and 0.5 μg/mL follicle stimulating hormone.
Frozen-thawed porcine ear skin fibroblasts (4-6 passages) were cultured in DMEM (Gibco) supplemented with 15% FBS and 1% P/S (Nalgene) at 39℃, 5% CO2 in air for 6-7 days until they reached confluence to synchronize the cell cycle at the G0/G1 phase. Cells were trypsinized with 0.05% trypsin-EDTA and centrifuged (500 ×g, 5 min) in HEPES-buffered TCM-199 supplemented with 0.78 mM NaHCO3, 0.14 mM penicillin G, 0.08 mM streptomycin and 3 mg/mL BSA (TCM-BSA). Cells were cultured in TCM-BSA containing 10 μg/mL phytohemagglutinin-P (PHA-P) for 10 min prior to SCNT.
SCNT was carried out in HEPES-buffered TCM-BSA containing 5 μg/mL cytochalasin B. The cumulus cells of the in vitro matured oocytes were removed by vortexing in PBS supplemented with 0.1% (w/v) hyaluronidase and 0.1% (w/v) PVA for 3 min. The oocytes were enucleated by removing the MII chromosome mass and the first polar body. Enucleation was confirmed by staining the oocytes with 1 μg/mL Hoechst 33342 at 39℃ for 15-20 min. About 90% of the oocytes were enucleated successfully (data not shown). Donor cells were injected into the perivitelline space of the enucleated recipient oocytes.
The reconstituted oocytes were manually aligned between two wire electrodes (1-mm apart) of a fusion chamber overlaid with 0.3 M mannitol solution containing 0.1 mM MgSO4, 0.05 mM CaCl2 and 0.5 mM HEPES (Duchefa Biochemie, Haarlem, the Netherlands). For the fusion and activation, two pulses of 1.25 kV/cm direct-current (DC) were applied for 30 μsec each using a BTX Electro Cell Manipulator 200 (BTX, San Diego, CA, USA). After the fusion/activation treatment, the reconstituted oocytes were placed in PZM-3 and checked for fusion.
For IVF, the in vitro-matured oocytes were separated from the surrounding cumulus cells by gentle pipetting. The oocytes were inseminated with frozen-thawed spermatozoa (3×105 spermatozoa/mL) in a 100-μL droplet of IVF medium (40 oocytes per droplet) at 39℃, 5% CO2 in air for 6 h. The IVF medium was based on modified Tris-buffered medium (mTBM) containing 113.1 mM NaCl, 3 mM KCl, 7.5 mM CaCl2.H2O, 20 mM Trizma, 5 mM pyruvic acid, 0.07 mM/mL streptomycin sulfate, 11 mM D-glucose, 0.17 mM penicillin-G and 0.7mM caffeine. After insemination culture, second polar body-extruded embryos were used for in vitro culture.
After activation or fertilization, the SCNT and IVF embryos were cultured in PZM-3 medium (Yoshioka et al., 2002) for 6 days at 39℃, 5% CO2 in air. At 3 h, 20 h and 6 days of culture, the embryos were sampled to analyze the x-box binding protein 1 (Xbp1) mRNA, ER stress-associated genes and apoptotic genes.
Total RNA from cells with or without TM treatment was extracted using Trizol (Invitrogen, Carlsbad, CA, USA). Poly(A) mRNAs of the IVF and SCNT embryos at 3 h and 24 h (one-cell stage) or 6 days (blastocyst stage) after IVF or fusion were extracted according to the manufacturer’s protocol using the Dynabeads mRNA Direct kit (Life Technologies AS, Oslo, Norway). Briefly, each group of embryos was sampled in 10 μL RNAlater® solution and cryopreserved at –70℃ until use. After thawing, 10 μL of Dynabeads oligo (dT) was added to each sample at room temperature and hybridized for 2 min, and lysed in 200 μL of lysis/binding buffer at room temperature for 2 min. The beads were separated from binding buffer using a Dynal magnetic bar. Poly(A) mRNAs and beads were washed with Buffer A and B and poly(A) mRNA was collected at 63℃ with 6.5 μL of Tris-HCl buffer. The cDNA synthesis was performed according to the manufacturer’s protocol using ReverTra Ace® qPCR RT Master Mix (Toyobo, Osaka, Japan). Six μL of the isolated total RNA or mRNA was denatured at 65℃ for 5 min. Genomic DNA was removed by adding 2 μL of 4x DN Master Mix to the RNA template and incubating at 4℃ for 5 min, followed by reverse transcription by adding 2 μL of 5x RT Master Mix and incubating at 37℃ for 20 min. Secondary RNA structure was denatured by incubating at 50℃ for 5 min and the reaction was terminated by incubation at 98℃ for 5 min. The products were stored at 4℃ until amplification.
Expression of Xbp1 mRNA, the key transcription factor in the ER stress condition, and apoptotic gene expression were detected by RT-PCR analysis, and expressions of ER stress-associated genes, the C/EBP homologous protein (CHOP), binding protein (BiP), activating transcription factor 4 (ATF4) and glucose-regulated protein 94 (GRP94) were analyzed by RT-qPCR. For RT-PCR, the cDNA samples were reverse-transcribed according to the manufacturer’s instructions using AccuPower® Taq PCR PreMix (Bioneer, Daejeon, Korea). After an initial denaturation step of 1 min at 72℃, 34 amplification cycles were performed. Each cycle consisted of denaturation at 95℃ for 30 sec, annealing at 58℃ for 30 sec, and extension at 72℃ for 50 sec. A final extension step of 5 min at 72℃ was performed to complete the reaction. The PCR products were analyzed by UV irradiation with an UV transilluminator (Bio-Rad, Berkeley, CA, USA) on a 2% agarose gel (Amresco, Cleveland, OH, USA) stained with ethidium bromide (Bioneer). Band intensity was measured by densitometry through ImageJ software 1.37v (National Institutes of Health, Bethesda, MD, USA). RT-qPCR was performed using the power SYBR Green PCR master Mix (TOPrealTM qPCR 2X PreMIX; SYBR Green with high ROX, Enzynomics, Daejeon, Korea) in a StepOne Plus instrument (Applied Biosystems, Foster City, CA, USA). The comparative CT method (∆∆CT method) was used for relative quantification of the mRNA level of each target gene. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as an internal control for the normalization of target gene expression. Primer pairs synthesized for the primer sequences of each gene are shown in Table 1.
RESULTS
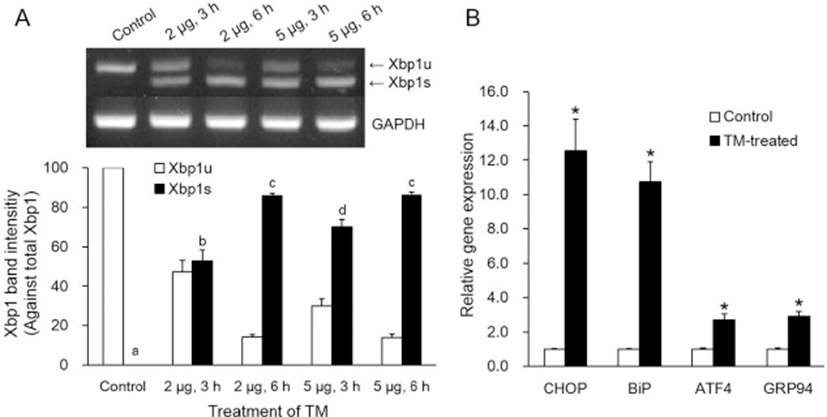
Only TM-treated cells, not untreated control cells, showed spliced Xbp1 (Xbp1s) mRNA (Fig. 1A). Xbp1s mRNA against total Xbp1 mRNA was increased in somatic cells treated with TM in relation to the TM dosage (2 or 5 μg/mL) and treatment time (3 or 6 h, p<0.05). ER stress-associated gene (CHOP, BiP ATF4 and GRP94) transcription levels were significantly higher after treatment with 5 μg/mL TM for 3 h compared with the corresponding control (p<0.05, Fig. 1B).
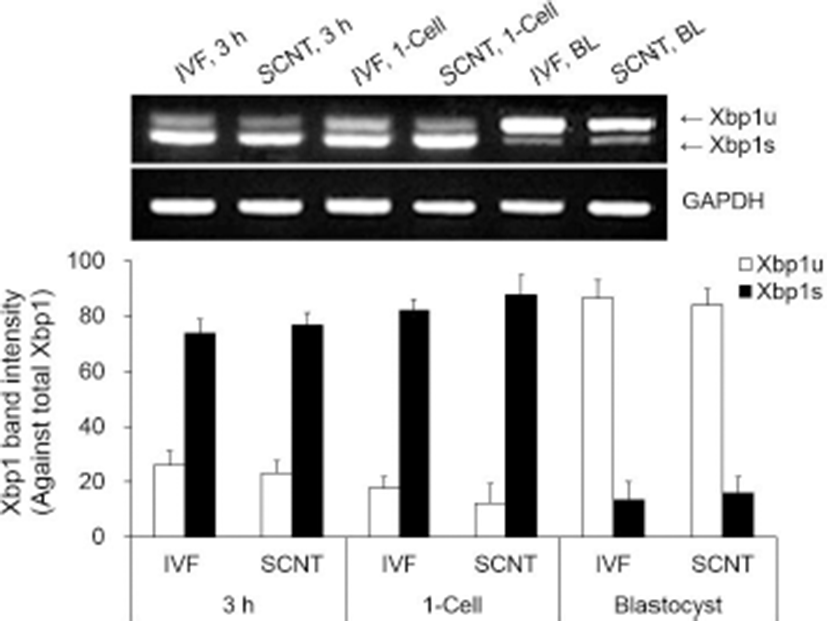
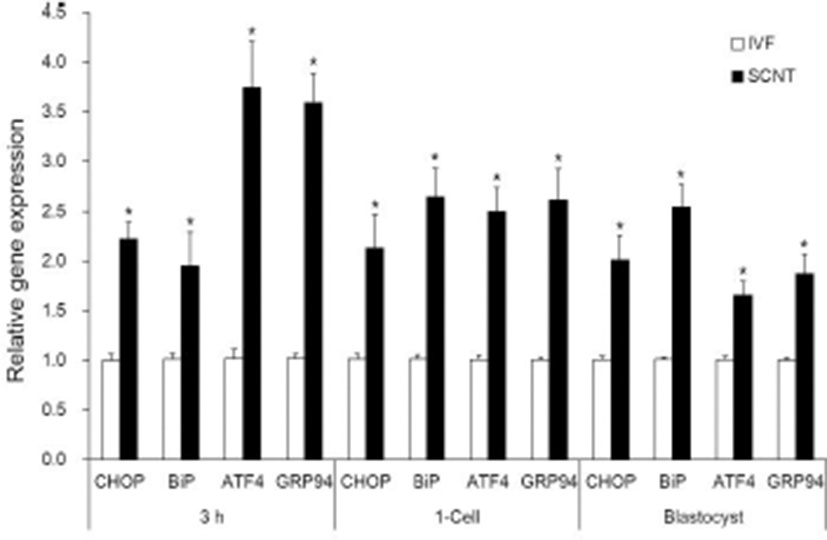
All groups of both IVF and SCNT embryos showed Xbp1s mRNA. Xbp1s mRNA expression was strong at 3 h after fusion/activation or IVF and the 1-cell stage, but weak in the blastocyst stage (Fig. 2, upper panel). Expression level of Xbp1s mRNA against total Xbp1 mRNA in the SCNT embryos was slightly higher in all the embryonic stages compared to IVF embryos, but there was no significant difference between SCNT and IVF embryos within the same stage (Fig. 2). However, ER stress-associated gene (CHOP, BiP ATF4 and GRP94) transcription levels were significantly higher in the SCNT embryos compared to IVF embryos in all the embryonic stages (p<0.05, Fig. 3).
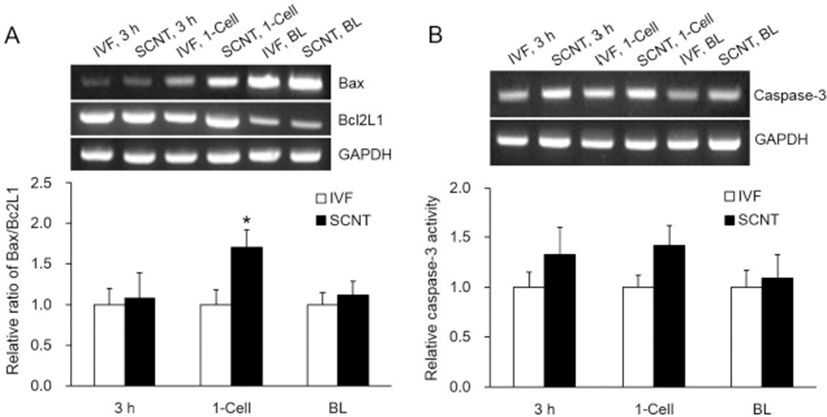
The ratio of Bax to Bcl2L1 was significantly higher in the 1-cell stage of the SCNT embryos (p<0.05), but no significant differences were observed in the other stages (Fig. 4A). Caspase-3 activity was greater in the SCNT embryos in the early stages, but there were no significant differences across the stages (Fig. 4B).
DISCUSSION
As unfolded/misfolded proteins continue to accumulate in the ER lumen, UPR is activated to adapt to the ER stress. UPR is important to restore ER homeostasis and reestablish normal ER function. To reduce the accumulation of unfolded/misfolded protein, the UPR signaling sensor/pathway inositol-requiring enzyme 1 (IRE1), PKR-like ER kinase (PERK) and activating transcription factor 6 (ATF6) are activated in the ER luminal domains (Boyce & Yuan, 2006; Yoshida, 2007). In general, BiP binds to the IRE1, PERK, and ATF6 and is inactive. As unfolded/misfolded proteins accumulate the ER, BiP dissociates from the IRE1, PERK, and ATF6 leading to the activation of UPR (Yoshida, 2007). Upon ER stress, IRE1 phosphorylation is activated through dimerization and unconventional splicing, the unspliced Xbp1 (Xbp1u) mRNA removing a 26-base nucleotide intron to cause a subsequent frame shift, and then converting to the Xbp1s (Schröder and Kaufman, 2005; Hetz and Glimcher, 2009). Activation of PERK involves homodimerization and autophosphorylation, leading to phosphorylation of α-subunit of eukaryotic translational initiation factor 2 (elf2α), which leads to the translation of ATF4 (Fels & Koumenis, 2006). ATF4 stimulates the expression of CHOP, a key transcription factor for initiating the apoptotic program in extreme ER stress conditions (Nishitoh, 2012). Activated ATF6 is moved to the Golgi apparatus for translocation (Yoshida, 2007). Translocated ATF6 regulates the expression of a number several genes including GRP78, GRP94, XBP1, and CHOP (Yoshida et al., 2001; Adachi et al., 2008; Parmar & Schröder, 2012). Successful UPR signaling results in embryo/cell survival, whereas the failure to restore ER homeostasis causes cell death.
We confirmed that porcine somatic cells treated with TM showed splicing of Xbp1 mRNA. TM is an antibiotic that inhibits N-glycosylation, which is often essential for protein folding and is usually used to induce ER stress in cultured cells (Dorner et al., 1990). Xbp1s mRNA was clearly detected by RT-PCR, and the expression level of Xbp1s revealed that the ER stress induced corresponded with TM dosage and treatment time. TM-mediated ER stress induction was also confirmed by ER stress-associated gene expression. These results are consistent with the ER stress analysis in embryos in this study. In previous studies, TM induced ER stress in cells and resulted in increased apoptosis (Boyce & Yuan, 2006; Zhang et al., 2014). Although, apoptosis did not significantly increase due to TM treatment in this study, ER stress-derived apoptosis could not be fully evaluated because we analyzed the apoptosis following ER stress after treatment with TM for a very short period only (data not shown). On the other hand, the CHOP pathway has been suggested a major regulator of ER stress-induced apoptosis (Zinszner et al., 1998). The strong expression of a pro-apoptotic CHOP, which called the growth arrest and DNA damage inducible gene 153 (GADD153, Price & Calderwood, 1992) revealed that apoptosis was induced by ER stress. Under the ER stress environment, CHOP stimulates a transcriptional profile that facilitates a pro-apoptotic program and also induces the death receptor 5 (DR5), which sensitizes cells to apoptotic stimulation and may activate Caspase cascades (Yamaguchi & Wang, 2004).
Xbp1s mRNA was stage-dependently detected at the various developmental stages of porcine parthenogenetic embryos showing different expression levels (Zhang et al., 2012b). ER stress-induced Xbp1 splicing has been implicated in the regulation of early porcine embryonic genome activation (Zhang et al., 2012b). Marginal ER stress may be essential for embryonic development but in excess may induce apoptosis and cell death (Yoshida, 2007). In this study, Xbp1s mRNA was successfully detected in the SCNT and IVF embryos at the early stage and blastocyst stage. Interestingly, there was no difference in Xbp1s mRNA expression levels between the SCNT and IVF embryos, although slightly increased Xbp1s mRNA levels against total Xbp1 mRNA were observed in SCNT embryos in this study. The quantification of Xbp1 mRNA expression level may not be accurate because we measured the band intensity of the RT-PCR products using densitometry. In a previous study, semiquantitative PCR method was used for the quantification of Xbp1 mRNA expression and the relative transcription levels of Xbp1s to total Xbp1 were significantly higher in the SCNT blastocysts compared with the IVF embryos (Song et al., 2014). Unlike Xbp1s mRNA expression, ER stress-associated genes, such as CHOP, ATF4, BiP, and GRP94, quantified by RT-qPCR were highly expressed in the SCNT embryos compared with the IVF embryos across all stages. ER stress-associated genes were highly expressed at 3 h immediately after SCNT and the 1-cell stage as well as in the blastocyst stage, suggesting that SCNT process induced ER stress in SCNT embryos.
It was demonstrated that ER stress-induced apoptosis was associated with the Caspase-3 (Groenendyk & Michalak, 2005) and Bcl-2 family protein (Szegezdi et al., 2006). Prolonged UPR induces expression of Caspase-3 and Bcl-2 family proteins resulting in apoptosis (Tabas & Ron, 2001; Gorman et al., 2012). In this study, the ratio of Bax to Bcl2L1 was higher only at the 1-cell stage of the SCNT embryos, and there was no difference in the Caspase-3 mRNA levels between the SCNT and IVF embryos. However, as in TM-treated somatic cells, sustained elevation of pro-apoptotic CHOP expression in the SCNT embryos revealed that apoptosis was aggravated by the SCNT process. On the other hand, in previous studies on porcine or bovine SCNT embryos, apoptosis was greater in the SCNT embryos compared with IVF or in vivo-derived blastocysts (Hao et al., 2003; Cánepa et al., 2014; Song et al., 2014). In our previous studies too, DNA fragmentation in the 1-cell stage of the SCNT embryos (Hwang et al., 2013; Bae et al., 2015) and apoptotic cells in the SCNT blastocysts by TUNEL assay were greater compared to IVF embryos (Bae et al., 2015).
In conclusion, the results of this study confirm that porcine SCNT process induced excessive ER stress and could lead to ER stress-induced apoptosis.