INTRODUCTION
Mammalian spermatogenesis is a complex developmental process which is dependent on a specific environment provided by the anatomical and cellular relationships in the seminiferous tubules (Oakberg, 1956). The whole process can be subdivided into three stages: (i) a pre-meiotic phase characterized by an increase in cell number due to mitotic division of diploid spermatogonia, (ii) a meiotic phase, which leads to the formation of haploid round spermatids and (iii) a post-meiotic phase which includes the morphogenetic events required for spermatozoa formation (spermiogenesis) (Griswold, 2016). Such multistep pathway in turn requires the invocation of numerous cellular processes and a strict regulation of cell specific gene expression in germ cells as well as surrounding somatic cells. However, the mechanism of spermatogenesis remains still elusive. To understand the mechanism of testicular germ cell differentiation, it is of great importance to isolate germ cell specific genes and characterize their functions. The polymerase chain reaction (PCR)-coupled subtractive hybridization technique called Representational Difference Analysis (RDA) have been used to identify testis-specific genes which are expressed only in adult rat testis and not in pre-pubertal testis (Lisitsyn & Wigler, 1993; Hubank & Schatz, 1994; Bowler, 2002; Gomes et al., 2005; Oh et al., 2013).
Molecular cochaperones which belong to family of Hsp40/ DnaJ have to be recruited by Hsp70 for its proper activity (Cyr et al., 1994, Kim et al., 2015). Hsp40/DnaJ, J family protein, is defined by the presence of a J domain (Walsh et al., 2004). A sequence based classification system has been established (Cheetham & Caplan, 1998; Ohtsuka & Hata, 2000). Type I DnaJ protein have all the three distinct domains: (i) a highly conserved J domain of approximately 70 amino acid residues in which the tripeptide histidine-proline-aspartic acid (HPD, J box) is located between the predicted helices II and III, (ii) a glycine and phenylalanine (G/F) rich region and (iii) a cysteine rich region resembling a zinc finger domain (Bork et al., 1992). Type II DnaJ proteins have a J domain linked by a G/F rich region to while type III DnaJ proteins have a J domain but lack the other sequence features that are found in type I and II members of the family.
As many as 41 different J family proteins have been identified to date (Meccariello et al., 2014). The first J family protein, MSJ-1, which was expressed specifically in male germ cell was reported (Berruti &Martegani, 1998) and followed by numerous reports on DnaJ family proteins which function during spermatogenesis as well as spermiogenesis (Meccariello et al., 2014). Type III DnaJ family protein, rDJL, was reported to be localized in acrosome region of spermatozoa (Yang et al., 2005). It has been reported that a mutant type I DnaJ protein (DjA1) can cause severe defect in spermatogenesis as well as aberrant androgen signaling in vivo (Terada et al., 2005). In the present study, we report the cloning of a type III member of DnaJ protein family, DnaJC18 cDNA and expression pattern of the gene at mRNA and protein levels in male germ cell of rat testis.
MATERIALS AND METHODS
Male Sprague Dawley rats and New Zealand rabbits were purchased from Daehan Biolink (Korea). RNAzol B reagent was purchased from Biotecx Laboratories (TX, USA). Nylon and nitrocellulose membranes were purchased from Sigma (MO, USA). Restriction enzymes were purchased from DCC-BIONET (Korea). The [α-32P] dCTP (3,000 μCi/ mmol) was purchased from Amersham Biosciences (England). DMEM, fetal bovine serum (FBS), trypsin-EDTA, antibiotics, and PBS were purchased from Invitrogen (MD, USA). The experiments using animals were conducted in accordance with the guidelines of Chonnam National University’s Animal Care and the National Research Council Guide for the Care and Use of Laboratory Animals.
Total RNA was isolated from 100-600 mg of tissue using the acid phenol guanidinium method (Chomczynski & Sacchi, 1987). Tissues were submerged in RNAzol B and homogenized with Biomixer (Nissie, Japan) for 1-2 min at 2,000 rpm. One-tenth volume of chloroform was added to the homogenate and the mixture was vortexed vigorously for 20 secs. The suspension was centrifuged for 15 mins at 14,000 rpm and the upper aqueous phase was taken. Equal volume of isopropanol was then added to the aqueous solution and the sample was kept for 15 mins on ice followed by centrifugation at 14,000 rpm for 15 mins. The RNA pellet was washed with 75% ethanol by centrifuging for 10 mins at 14,000 rpm. All centrifugations were done at 4℃. Finally, the pellet was dried for 10 mins, resuspended in DEPC treated water by vortexing and quantified by optical density measurement.
In order to identify the specific genes expressed only in adult rat testis, subtracted cDNA library was constructed using Representational Difference Analysis (RDA) [Hubank & Schatz, 1994; Gomes et al., 2005; Oh et al., 2013]. RDA was carried out with a commercial kit (PCR select, BD Clontech, CA, USA). Driver cDNA population (D-cDNA) and tester cDNA population (T-cDNA) were prepared from prepubertal (2-week-old) and adult (8-week-old) male rat testis, respectively. Briefly, 2 μg of cDNAs were digested with RsaI and the digested T-cDNAs were then divided into two fractions and ligated to two different primers, respectively, with T4 DNA ligase. First hybridization was then carried out by adding excess D-cDNA to each T-cDNA fraction in the thermal cycler at 98℃ for 1.5 mins and at 68℃ for 8 h. This was followed immediately by second hybridization with fresh denatured D-cDNA at 68℃ overnight. Multiple PCR reactions were set up to generate the initial amplicons (representations). Each 25 μL reaction contained 1 μL diluted subtracted sample, 1x PCR buffer, 10 mM dNTP mix, 10 μM PCR primers and Advantage cDNA polymerase mix (BD Clontech 8430-1, CA, USA). After the 3’ ends of the adapters were filled, twenty-seven cycles of amplification were then performed at 94℃ for 10 secs, 66℃ for 30 secs and 72℃ for 1.5 mins. During amplification, cDNAs presumably from the adult testis were to be exponentially amplified. The final PCR products were resuspended in 1x TE at about 0.5 μg/μL.
One of the clone, 230 bp fragment (tsg-107) from the above subtracted library was used to screen rat testis cDNA library (BD Clontech, RL300a, CA, USA) to obtain full-length cDNA clones (Sambrook & Russel, 2001). In short, about half a million plaques were plated on ten top LB agarose plates with a density of about 5×105 plaques per plate. When the diameter of the plaques reached around 0.5-0.7 mm, the phage particles were transferred on to a nylon membrane followed by denaturation and neutralization. The DNA was then cross-linked to the membranes by UV. The membranes were incubated in prehybridization solution (6 X saline-sodium-citrate, SSC; 0.1% SDS; 5 X Denhardt solution; 100 μg/mL of denatured Salmon sperm DNA) at 65℃ for 2 h. Then, [α-32P]-labeled 230 bp cDNA was added to the fresh prehybridization solution (1×106 cpm/mL) and incubated at 65℃ for 20 h. The random priming procedure was used to radiolabel the DNA (Feinberg & Vogelstein, 1983). The membranes were washed with 2×SSC/0.1% SDS at RT for 30 mins twice followed by a final wash with 0.3 × SSC/0.1% SDS at 50℃ for 30 mins twice and were exposed to X-ray film at –80℃ for autoradiography.
Total RNA (about 10 µg) from each sample was fractionated in a 1% formaldehyde/agarose gel and transferred on to a nylon membrane followed by UV cross-linking (Lee et al., 1997). Northern blots were pre-incubated in a solution containing 50% formamide, 50 mM NaH2PO4, 5× Denhardt solution, 5 × SSC, 0.1% SDS, 2% Dextran, 1 mM EDTA and 100 μg/mL of denatured salmon sperm DNA at 42℃ for 12 h. Hybridization was performed with the [α-32P]-labeled cDNA probe (pCG1.3) prepared by random priming procedure as described previously under the same conditions of pre-incubation for 24 h. Following hybridization, blots were washed in 2 × SSC/0.1% SDS for 10 min at room temperature; 1×SSC/0.1% SDS for 30 min at 65℃, and 0.1×SSC/0.1% SDS for 30 min at 65℃. After washing, the membrane was exposed to X-ray film for 24 h for autoradiographic signals.
The in situ hybridization was carried out as described (Lee et al., 1999). Adult rat testis was fixed at 4℃ for 6 h in 4% paraformaldehyde in PBS followed by overnight immersion in PBS containing 0.5 M sucrose. Cryostat (Leica, Germany) sections of the testis (14 μm thick) were mounted onto microscope slides coated with poly-L-lysine, fixed in 4% paraformaldehyde solution and stored at –80℃ until analysis. Sections were pretreated serially with 0.2 M HCl, 2 × SSC, pronase (0.13 mg/ml), paraformaldehyde and acetic anhydride in triethanolamine. Hybridization was carried out at 52-55℃ overnight in the mixture containing [35S]-labeled pCG1.3 insert as a probe (108 cpm/mL), 50% formamide, 0.3 M NaCl, 10 mM Tris-HCl, 5 mM EDTA, 1 × Denhardts solution, 10% Dextran Sulfate, 1 μg/mL carrier transfer RNA and 10 mM dithiothreitol. Post hybridization washing procedure was performed under stringent conditions, which included ribonuclease A (25 μg/mL) treatment at 37℃ for 30 mins and a final stringency of 0.1 × SSC. Slides were dipped into NTB-2 emulsion (Eastman Kodak Co., NY, USA) and exposed at 4℃ for 2 weeks until development. The slides were stained with hematoxylin, counterstained with eosin and examined under a light microscope with bright and dark field illumination.
The BSA conjugated with 20 amino acids peptide –NH2 GLYRDERLRQKAESLKLENC COOH-, corresponding to residues #325-#344 of DnaJC18 protein was synthesized by AnyGen Co. Ltd. (Korea). The BSA conjugated synthetic peptide was dissolved at the concentration of 0.2 mg/mL in physiological saline and emulsified with 1 ml of Freund complete or incomplete adjuvants. Two New Zealand rabbits were immunized with 0.2 mg of fusion protein per single immunization, four times at intervals of two weeks. After three booster injections of antigen, blood was collected. The affinity purification of the antibody was carried out with Melon Gel IgG purification kit (Pierce, IL, USA) and subsequently used for western blot analysis and immunohistochemistry.
The rat tissue homogenates from 8-week-old rats were fractionated as described (Laemmli, 1970) and subjected to 10% SDS-PAGE. Proteins were electroblotted to nitrocellulose membranes (Sigma, MO, USA) using a Trans Blot Cell (Bio-Rad Laboratories, CA, USA). Membranes were hybridized with rabbit anti-DnaJC18 (1:500 dilution), α-tubulin (1:2000 dilution) antibodies (Sigma, MO, USA) or preimmune serum. Primary antibodies were detected using anti-rabbit IgG (1:3000 dilution) conjugated with Horse Radish Peroxidase (HRP) (Sigma, MO, USA). Reactive protein bands were visualized by enhanced chemiluminescence (ECL) using the manufacturer's protocol (Amersham Pharmacia, UK).
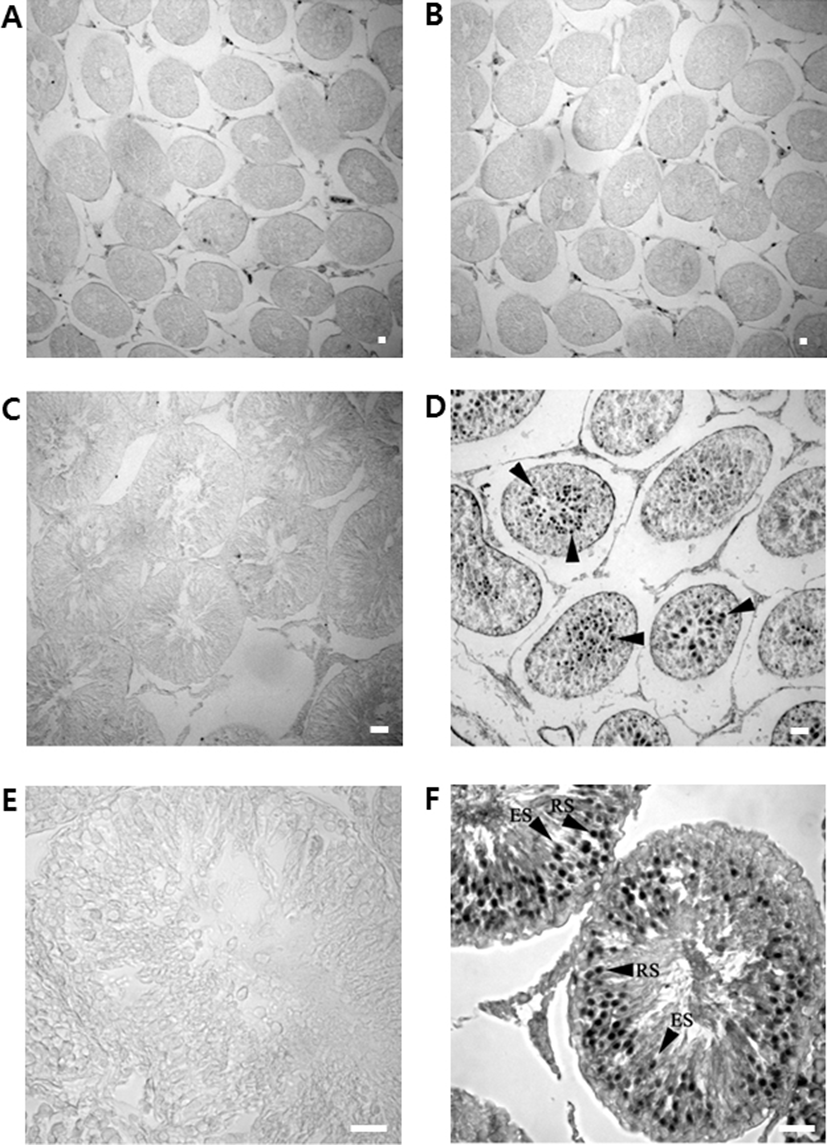
Testicular tissues from 3, 5 and 9-week-old rats were post- fixed in 4% neutral buffered paraformaldehyde (Sigma, MO, USA) solution and embedded in paraffin according to standard procedure (Gomes et al., 2005). Testicular cross sections (6 µm thick) were dewaxed in two changes of xylene for 10 min and rehydrated through a series of alcohol to the single-strength PBS. Sections were then incubated in 3% H2O2 (in methanol) for 30 min at room temperature and rinsed in PBS three times (for 5 mins each). A blocking step was performed by placing slides in 2% normal rabbit serum (NRS) in PBS for 1 h and then rinsing in PBS three times (for 5 mins each). Sections were incubated overnight at 4℃ with rabbit anti-DnaJC18 antibodies diluted (1:300) in blocking solution. Slides were rinsed in PBS and incubated with rabbit anti-goat IgG (Vector Laboratories, CA, USA) for 1 h at room temperature. The slides were rinsed in PBS and incubated with ABC reagent (Vecta-stain Kit; Vector Laboratories, CA, USA) for 1 hr. Finally, the slides were incubated with diaminobenzidine (Sigma, MO, USA) until a brown color developed. The color reaction for the control (with preimmune rabbit serum) slide was terminated at the same time as for the slides treated with primary antibody. Slides were cleared with xylene and mounted under Permount (Fisher Scientific, PA, USA). Images were taken using the Image Pro Express software (MagnaFire SP imaging, Sciscope Instrument Company).
The plasmid pBSK(+)-DnaJC18 harboring the cDNA of rat DnaJC18 was used as the PCR template to construct the GFP-DnajC18 fusion protein plasmid. The entire coding region of rat DnaJC18 cDNA was amplified with a forward primer containing the added EcoRI site at the upstream of the start codon (5’ TAGAATTCTATGGCGGCCACTCTG GGC 3’) and a reverse primer including the SalI site at downstream of stop codon (5’ TAGTCGACTCAGCCGGC CCTGCGGAG 3’). The PCR products were digested with EcoRI/ SalI and inserted in frame into the EcoRI and SalI site of pEGFP-C1 mammalian expression vector (BD Clontech, CA, USA). The GFP-DnaJC18 constructs (pEGFP-DJC-5) was confirmed by DNA sequencing.
CVI (kidney cell line) and GC2 (germ cell line) cells were maintained in DMEM medium supplemented with 10% fetal bovine serum and used for the transfection experiments and. Cells were seeded in Lab-Tek II chamber slide (Nalge Nunc Inc., NY, USA) with 2 ×104 cells per chamber in 150 µL of medium. Cells were transiently transfected with 0.2 µg of each pEGFP-C1 vector (control plasmid) and pEGFP-DJC-5 (GFP-DnaJC18 fusion construct) using Lipofectamine Plus reagent (Qiagen, CA, USA) according to the manufacturer’s instruction. The transfected cells were subjected to localization study with confocal microscopy after 24 hours of transfection. A Zeiss LSM 410 confocal microscope equipped with an external krypton/argon laser was used for localization studies. Image processing was performed using IP Labs (Scanalytics) and Adobe Photoshop 7.0 (Park & Cheon, 2015).
RESULTS
About one hundred clones from the subtracted cDNA library were selected randomly. In order to confirm whether the selected clones are testis-specific novel genes or not, preliminary rat tissue Northern blot analysis was performed with the inserts derived from the cDNA clones (data not shown). The preliminary Northern blot analysis result revealed that approximately 30% clones of subtracted cDNA library showed the testes specificity. One clone (tsg-107, insert size 230 bp) was sequenced followed by Blastx on the GenBank database. The result revealed that it showed significant homology with Hsp40/DnaJ protein family at the amino acid level. Therefore, this clone was further studied.
To obtain the full-length cDNA clone the rat testis cDNA library was screened with cDNA fragment of tsg-107 as a probe as described in Materials and Methods. One clone, λCG 1.3, was isolated and then the insert was subcloned into pBSK(+) vector, pCG 1.3. The clone pCG 1.3 contained the 1.4kb cDNA fragment. The sequencing the insert of pCG 1.3 clone showed that it has 1438 nucleotide long insert and generated the longest open reading frame (ORF) of 357 amino acids from nucleotide #178 to #1250 as shown in Fig. 1. (GenBank Accession No. DQ158861). The deduced protein sequence is found to contain the J domain (grey background, aa #81 to #138, Fig. 1) along with the J box (open box, HPD, Fig. 1) without any additional signaling domain such as G/F rich region or C rich domain. Therefore, it is regarded as a typical member of type III subfamily of HSP40/DnaJ protein family.

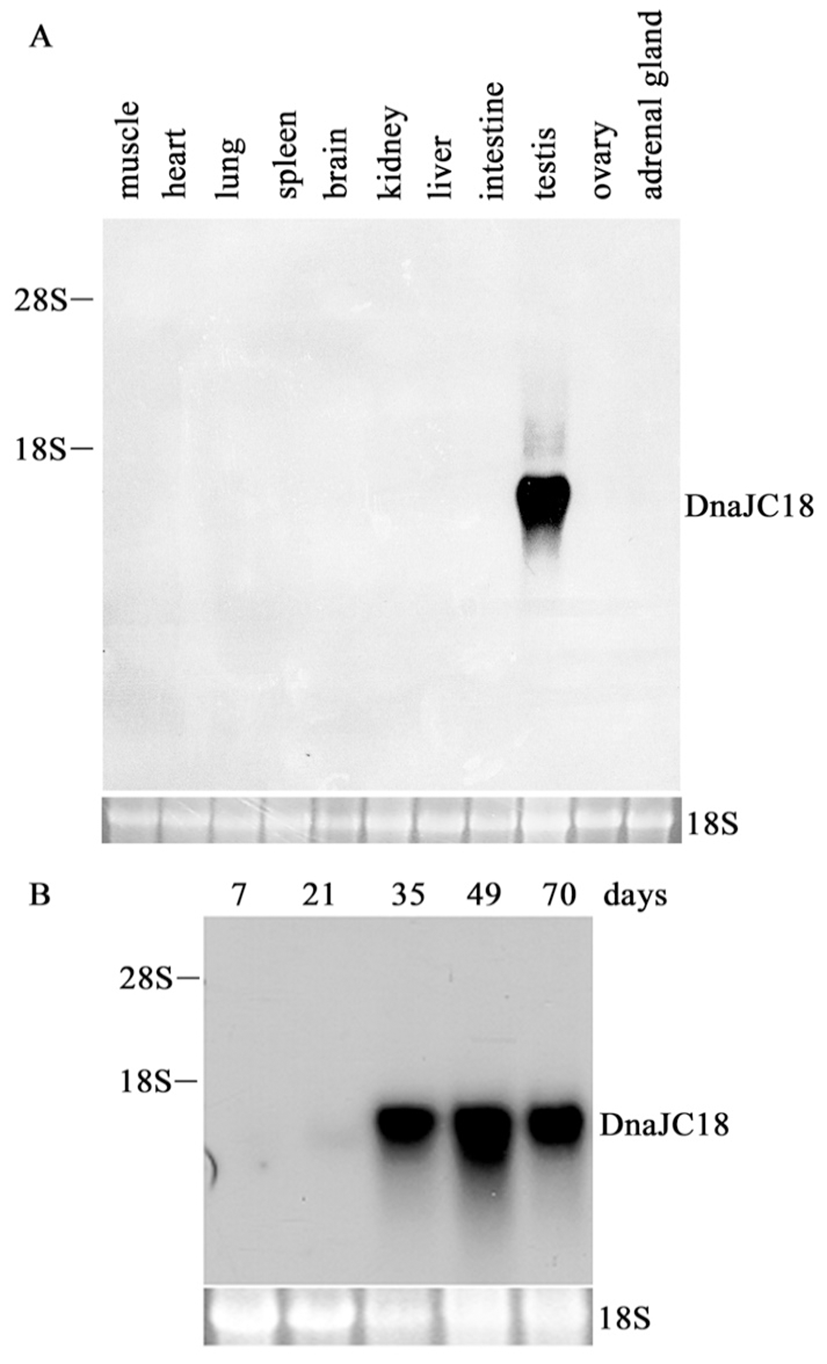
To investigate the tissue expression pattern of rat DnaJC18 mRNA, rat multiple tissue blot was hybridized with insert from pCG1.3 as a probe. A transcript whose size is believed to be about 1.4 kb was exclusively found only in testis (Fig. 2A), indicating that DnaJC18 gene expression is restricted in mature testis among tissues tested. Total RNA extracted from testis of 7, 21, 35, 49 and 70 day-old rats were subjected to Northern blot analysis. There was a faint signal after 21 days of testis, and strong signals were detected from postnatal day 35 and expressed up to adulthood (Fig. 2B). These data indicated that the expression of the DnaJC18 gene was expressed strongly only in postpubertal rat testis.
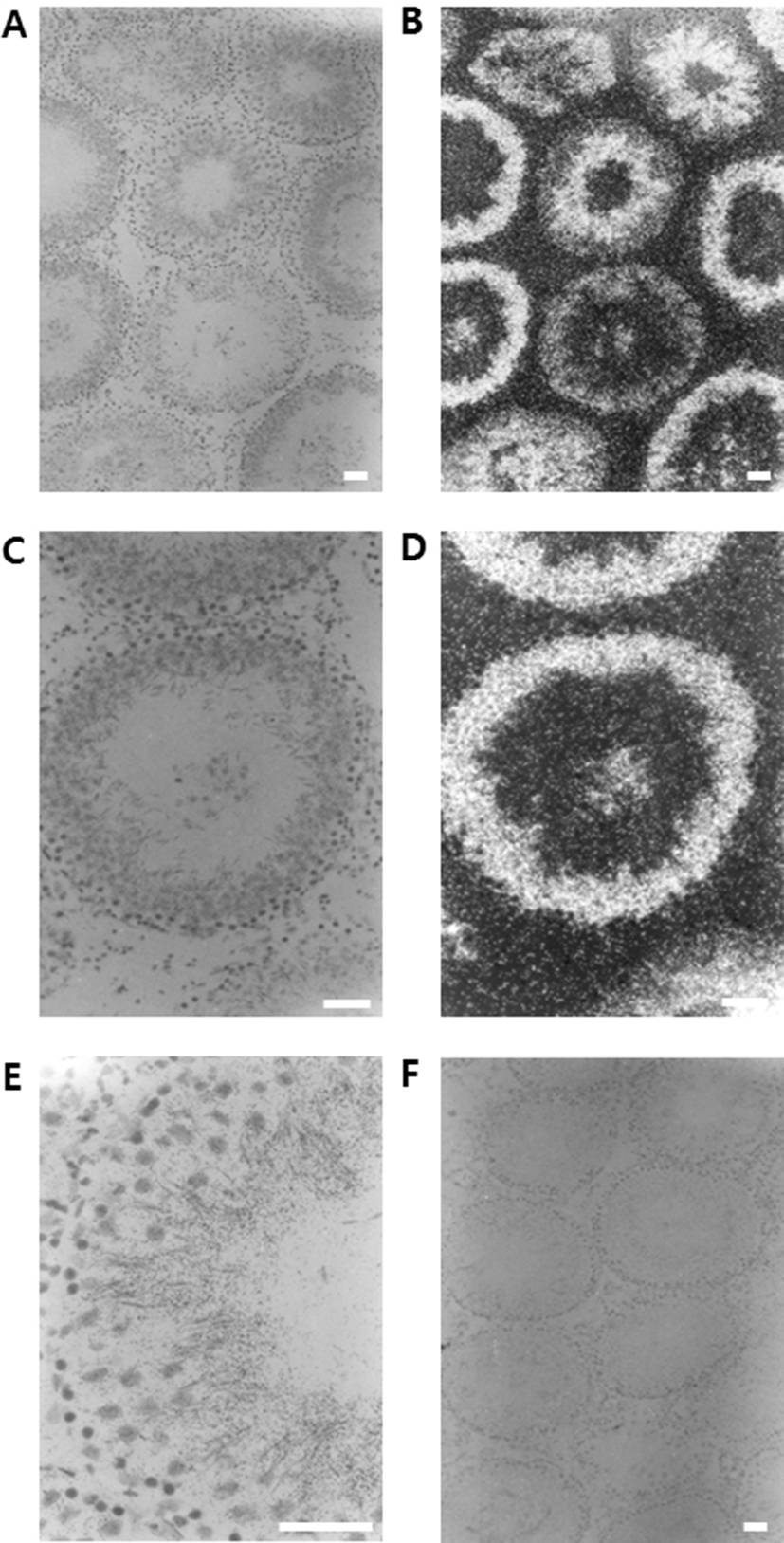
In order to check the cellular distribution of DnaJC18 mRNA in the adult rat testis, in situ hybridization was carried out. The positive signals were detected exclusively in the developing germ cells. Moreover, the expression level of DnaJC18 mRNA was detected from mid pachytene to early elongated spermatids, with its maximum expression in the round spermatids (Fig. 3E). The signals were detected as dark and white spots in the bright (Fig. 3A and 3C) and dark fields (Fig. 3B and 3D) of photoradiographs, respectively. Negative control section hybridized with a sense DnaJC18 probe showed no signals (Fig. 3F).
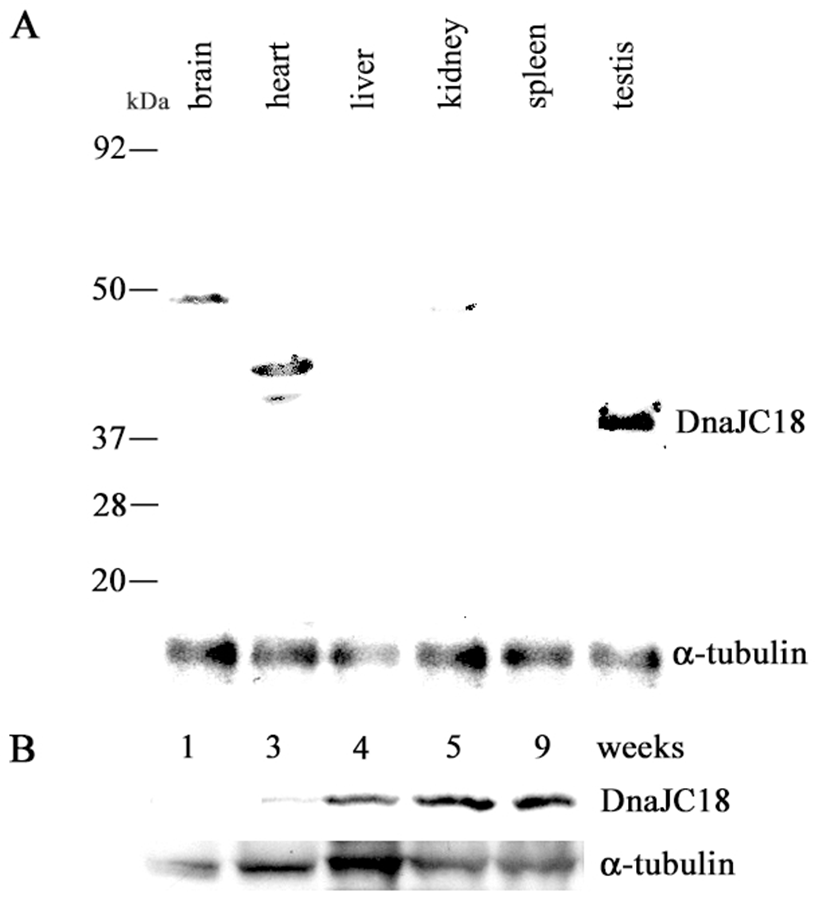
Rabbit polyclonal DnaJC18 antibody was raised against the synthetic peptide as described in Materials and Methods. The antibodies were affinity purified and the specificity of the antibodies was confirmed by Western blot analysis using in vitro translated DnaJC18 protein (data not shown). The 1071 bp long ORF of DnaJC18 having 357 amino acid residues was predicted to produce a protein with a molecular weight of about 41.2 kDa. A band of approximately 41.2 kDa size of protein was detected only in the testis (Fig. 4A). However nonspecific bands of higher molecular weight were observed in brain and heart as well. Western blot analysis was also carried out to check the expression pattern of DnaJC18 at various developmental stages of rat testes. There was faint expression of DnaJC18 protein in 3 weeks postnatal testis, which is consistent with Northern blot analysis. Significant level of DnaJC18 protein was expressed from testis of postnatal week 4 (Fig. 4B).
Sections of various developmental stages of rat testes were immunostained with the rabbit polyclonal anti-DnaJC18 antibodies. No positive signals were detected in germ cells of 3-week of prepubertal testis (Fig. 5B) even though faint bands have appeared on Northern blot and Western blot analysis in 3 weeks old testis. Strong immunostaining signals appeared from testis of postnatal week 5 (Fig. 5D) and postnatal week 9 (Fig. 5F). Adult testicular sections showed strong signals in well-differentiated seminiferous tubules mainly in round spermatids and to some extent in elongated spermatids (Fig. 5F).
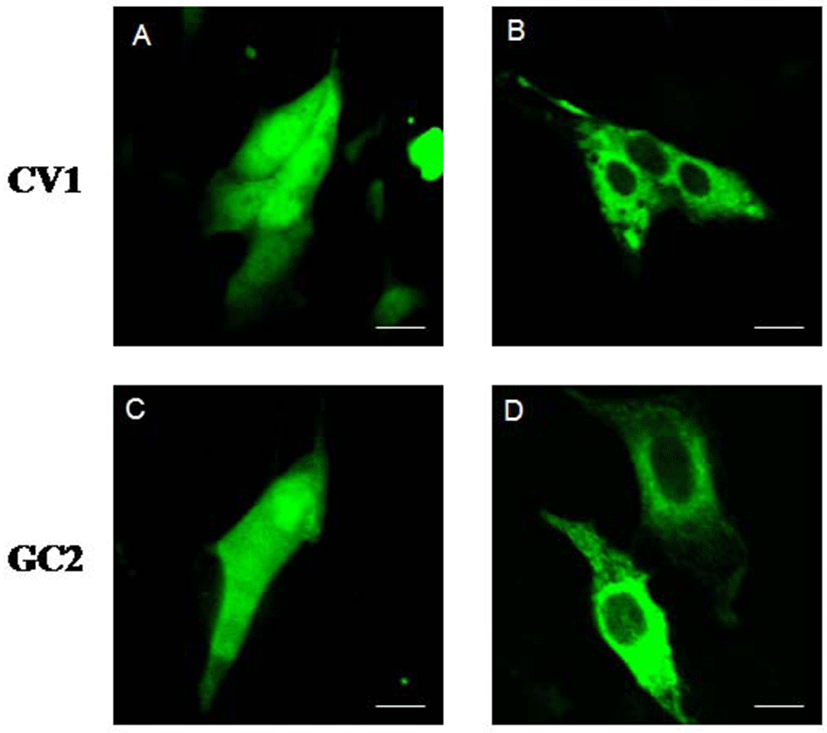
To ascertain the subcellular localization of type III DnaJC18 protein, we transfected CV1 (kidney cell line) and GC2 (germ cell line) cells with the plasmid encoding green fluorescent protein (GFP) and with GFP-DnaJC18 fusion protein, respectively, and carried out confocal microscopic studies. Transfection with pEGFP-C1 (control plasmid, GFP alone) showed ubiquitous expression of GFP protein in CV1 (Fig. 6A) and GC2 (Fig. 6C) cells. However, transfection of pEGFP-DJC-5 (GFP-DnaJC18 fusion construct) to both CVI (Fig. 6B) and GC2 (Fig. 6D) cells revealed that green fluorescence signals were found exclusively in the cytoplasm, suggesting that DnajC18 protein might be cytosolic protein.
DISCUSSIONS
Previously two testes specific type II DnaJ proteins MSJ-1 (Berruti et al., 1998) and MFSJ1 (Yu & Takenaka, 2003) were reported. J domain within DnaJ protein family is known to mediate interaction with the Hsp70 and regulate its ATPase activity (Meccariello et al., 2014). The difference between type III DnaJ family, DnaJC18, and type II DnaJ family, MSJ-1, and MFSJ1, resides in the amino acid sequence in the C-terminus of the protein. Unlike two testes specific type II DnaJ family member mentioned above, DnaJC18 does not contain the cysteine rich region. The cysteine rich region is the third canonical domain in DnaJ family proteins that shows structural homology with the zinc finger domain. This suggests DnaJC18 might not be a DNA binding protein and this was further confirmed by the observation that DnaJC18 was located in the cytoplasm of cells (Fig. 6).
It is conceivable that DnaJC18 protein might act together with the Hsp70 protein as it has been shown that one Hsp70 protein might form a complex with several different Hsp40/ DnaJ proteins. However, type III DnaJ family proteins is thought to be functionally different than type I and type II subfamily members since no type III DnaJ proteins has been shown to bind to Hsp70 protein as a cochaperone to date. Therefore, it is unlikely that these proteins function as molecular cochaperones.
Some type III DnaJ proteins assist the recruitment of a selective isoform of Hsp70 to a discrete site. In other cases, the Hsp70 and typeIII DnaJ protein are recruited independently to the site of actions where type III DnaJ protein productively stimulates ATP hydrolysis of its Hsp70 partner (Berruti & Marteganu, 2001).
The male germ cell specific expression of DnaJC18 at the mRNA transcript level as well as the protein level indicates that DnaJC18 might play a pivotal role in the process of spermatogenesis in adult testis. Further functional studies are needed to elucidate its role during rat male germ cell development.