INTRODUCTION
In vertebrates, facial skeletons arise in the cranial neural crest cells (cNCCs) that populate a series of pharyngeal arches. Besides cNCCs, the arches are composed of the pharyngeal endoderm (PE), head mesoderm, and pharyngeal ectoderm. Interactions among these tissues are crucial for craniofacial development, including the facial skeletons (Graham, 2008). In particular, PE is essential for cNCCs in the arches to form facial skeletal elements, as evidenced by the loss of facial skeletons in the absence of the endoderm of sox32 mutants in zebrafish (Piotrowski & Nüsslein-Volhard, 2000; Dickmeis et al., 2001; Kikuchi et al., 2001). During craniofacial development, PE forms a series of pharyngeal pouches, outpocketings of the PE, that segments the pharyngeal arches then provides signals, such as sonic Hedgehog and Jagged, for the cNCCs to differentiate into the facial skeletal elements (Miller et al., 2000; Piotrowski & Nüsslein-Volhard, 2000; Zuniga et al., 2010). In addition, the pouches continue to develop into endocrine glands in the head and neck, including the thymus and parathyroid (Grevellec & Tucker, 2010). Abnormal pouch formation has been associated with defects in facial skeletons, along with malformation or absence of the thymus and parathyroid in all vertebrates examined so far and is an etiology of DiGeorge syndrome (DGS), a congenital craniofacial disability, in human (Driscoll et al., 1992; Piotrowski & Nüsslein-Volhard, 2000; Lindsay et al., 2001; Tran et al., 2011). Despite the importance of pouches in craniofacial development, the genetic and cellular mechanisms underlying pouch formation are just being revealed. Tbx1 is a master regulator for developing pouches in fish to human, and TBX1 haploinsufficiency is the genetic cause of DGS (Lindsay et al., 2001; Piotrowski et al., 2003; Tran et al., 2011). Pax1/9 and Foxi1/3 are required for pouch formation in fish and mice (Peters et al., 1998; Nissen et al., 2003; Solomon et al., 2003; Edlund et al., 2014; Okada et al., 2016; Jin et al., 2018; Liu et al., 2020). Signaling molecules such as Fgf3/8, Wnt11r/4a, EphrinB2/B3, and Bmp are also necessary for pouch development in either zebrafish or mice (Crump et al., 2004; Herzog et al., 2004; Agrawal et al., 2012; Choe et al., 2013; Choe & Crump, 2015; Lovely et al., 2016; Li et al., 2019). Tbx1 regulates pouch morphogenesis by controlling Wnt11r and Fgf8a expression in the head mesoderm in zebrafish (Choe & Crump, 2014). Pax1 promotes pouch formation by expressing tbx1 and fgf3 in the PE in medaka and zebrafish, with Foxi1 controlling pouch development through ectodermal Wnt4a expression in zebrafish (Okada et al., 2016; Jin et al., 2018; Liu et al., 2020). The cellular process of pouch formation has been studied in detail in zebrafish. Pouch formation begins with destabilizing the bi-layered epithelial structure of the PE that is promoted by the Tbx1-Wnt11r pathway (Choe et al., 2013; Choe & Crump, 2014). In the destabilized PE, a population of endodermal cells, called pouch-forming cells, migrates out collectively toward the head mesoderm located adjacent to the PE, with Fgf8a expression in the head mesoderm acting as a chemoattractant (Choe et al., 2013; Choe & Crump, 2014). The migrating pouch-forming cells are rearranged into a bi-layered structure followed by re-stabilization of the pouch epithelia to complete pouch formation, with Wnt4a and EphrinB2/B3 required for the rearrangement and restabilization, respectively (Choe et al., 2013; Choe & Crump, 2015). Thus, at the cellular level, the migration and rearrangement of pouch-forming cells that cytoskeleton dynamics might control are important driving forces for pouch formation. Indeed, inhibiting actin cytoskeleton dynamics in chick caused severe defects in pouch formation, indicating an essential role for actin cytoskeleton dynamics in pouch development (Quinlan et al., 2004). However, any regulators of actin cytoskeleton dynamics required for pouch formation have not been identified yet in any vertebrates. Here, we analyze the expression and function of cofilin1-like (cfl1l), a regulator of actin cytoskeleton dynamics, during pouch formation in zebrafish.
The Cofilin family is included in the Actin-depolymerizing factor (ADF)/Cofilin superfamily that contains the conserved ADF homology (ADF-H) domain from yeast to human (Lappalainen et al., 1998; Ono, 2007). Traditional Cofilins (Cfls) consist of non-muscle type Cfl1, muscle type Cfl2, and ADF (also named Destrin) in mammals (Ono, 2007; Ohashi, 2015). In vertebrate cells, Cfls remodel the actin cytoskeleton by promoting depolymerization or polymerization of actin filaments, with the activity of Cfls being regulated by the cellular microenvironments such as pH, phosphatidylinositols, protein kinases, and phosphatases (Ono, 2007; Shishkin et al., 2016). For example, in vertebrate cells, while LIM-kinases and testicular protein kinases phosphorylate and inactivate Cfl1, slingshot protein phosphatases, protein phosphatases 1 and 2A, and chronophin dephosphorylate and activate Cfl1 (Ono, 2007; Shishkin et al., 2016). Consequently, vertebrate cells change their shape, migrate, and are rearranged through reactions of the phosphorylation/ dephosphorylation of Cfls, which are essential for developmental processes (Abe et al., 1996; Obinata et al., 1997). Indeed, Cfl1 knockout mice were embryonic lethal, with significant defects in neural crest cell migration and neural tube closure (Gurniak et al., 2005). Muscle-specific Cfl2 knockout mice were normal at birth. Still, they died at the postnatal stage due to the degeneration of myofibers, sarcomeric disruptions, and actin accumulation, indicating an essential role of Cfl2 in muscle maintenance (Agrawal et al., 2012). Cfl1 and Adf are also necessary for developing kidney and eye in mice. Kidney-specific deletion of Cfl1 in Adf mutants resulted in complete loss of kidney, with kidney-specific deletion of Cfl1 in the heterozygotic mutant for Adf resulting in the hypoplastic kidney (Kuure et al., 2010). A missense mutation in Adf caused defects in eyes, with the corneal epithelium abnormally thickening in mice (Ikeda et al., 2003). In zebrafish, cfl1 mutants showed defects in the heart and kidney, indicating a role for Cfl1 in heart and kidney development (Ashworth et al., 2010; Fukuda et al., 2019). In addition, a morpholino-mediated knockdown of cfl1 resulted in defects in morphogenic movements of the deep cell layer during gastrulation (Lin et al., 2010). Zebrafish contains three paralogs of the cfl gene, including cfl1, cfl2, and cfl1l, but no adf, in the genome, with the role of cfl2 and cfl1l during embryonic development undetermined. Here, we report the expression of cfl11 in the pharyngeal pouches, which is dispensable for head development, including pouch formation in zebrafish.
MATERIALS AND METHODS
According to the Animal Protection Act (2017) in Korea, zebrafish were grown and maintained. All zebrafish work was approved by Gyeongsang National University Institutional Animal Care and Use Committee. cfl1l mutant lines were generated by CRISPR/Cas9 system (Hwang et al., 2013). 150 pg of gRNA targeting the second exon of the cfl1l gene and 150 pg of mRNA encoding a nuclear-localized Cas9 were injected together into one-cell stage wild-type Tübingen (TU) embryos harboring Tg(~3.4her5:EGFP), a PE reporter transgene (Tallafuß & Bally-Cuif, 2003). Injected embryos were grown to adulthood and outbred to wild-type TU animals to identify zebrafish bearing in/del mutations in the cfl1l gene. Two cfl1l mutant lines (cfl1lGNU38 and cfl1lGNU39) were secured. For genotyping of cfl1lGNU38 and cfl1lGNU39, a PCR amplified cfl1l fragment with primers cflf1_GT_ F (5’-CCAAAGAGTGTCGCTACG-3’) and cfl1l_ GT_R (5’-CCCATCTGACAACGCTAC-3’), was digested with Sau96I, with a wild-type fragment producing 185 and 63 bp and mutant fragment generating 248 bp.
Phylogenetic analysis was carried out with the amino acid sequences of vertebrate homologs of Cfl and Adf proteins obtained from National Center for Biotechnology Information (NCBI): (Aa-Cfl1, Anguilla anguilla (European eel) XP_035266577; Om-Cfl1, Oryzias melastigma (Indian medaka) XP_024139063; Dr-Cfl1, Danio rerio (zebrafish) NP_998806; Xt-Cfl1, Xenopus tropicalis (tropical clawed frog) NP_998878; Mm-Cfl1, Mus musculus (house mouse) NP_031713; Hs-CFL1, Homo sapiens (human) NP_005498; Cm-Cfl2, Callorhinchus milii (elephant shark) XP_007891564; Aa-Cfl2, XP_035275350; Om-Cfl2, XP_024123733; Dr-Cfl2, NP_991263; Xt-Cfl2, NP_001011156; Mm-Cfl2, NP_031714; Hs-CFL2, NP_001230574; Aa-Cfl1l, XP_035255492; Om-Cfl1l, XP_024153813; Dr-Cfl1l, NP_998804; Cm-Cfl2l, NP_001280090; Xt-Adf, XP_002937490; Mm-Adf, NP_062745; Hs-ADF, NP_006861). Whole amino acid sequences deduced from the coding sequence (CDS) of vertebrate cfl and adf genes were aligned by CLUSTAL Omega with default parameters (Sievers & Higgins, 2021). The RAxML (Randomized Axelerated Maximum Likelihood) method with default parameters was applied to construct a maximum likelihood tree (Stamatakis, 2006).
Fluorescent in situ hybridization in conjunction with GFP immunohistochemistry (Torrey Pines Biolabs, 1:1,000), immunohistochemistry for Alcama (Zebrafish International Resource Center, 1:400), and Alcian Blue staining were carried out as described previously (Crump et al., 2004; Zuniga et al., 2011). Partial cDNA fragments of cfl1l were amplified from mixed-stage embryos with primers cfl1l_rF (5’-CCTCAGGTGTAGCGATCA-3’) and cfl1l_rR (5’-CCACCAAGTTTTTCCACA-3’). The PCR fragments were cloned into the pGEM®-T easy vector (Promega). Antisense riboprobes were transcribed with SP6 RNA polymerase (Roche Life Sciences) using digoxigenin (DIG)-labeled nucleotides (Roche) from sequence-verified plasmids as described previously (Jeon et al., 2019).
Fluorescent images were taken on an Olympus FLUOVIEW FV3000 confocal microscope using FV31S-SW software (Olympus). Approximately 100 μm Z-stacks at 3.5-μm intervals were captured with an Olympus UPLXAPO 20X objective lens, then were stacked into a single image. Facial cartilages were dissected manually from Alcian Blue staining larvae and were flat-mounted for imaging with an Olympus BX50 upright microscope using mosaic V2.1 software (Tucsen Photonics). Any adjustments were applied to all panels using Adobe Photoshop (Adobe Systems).
RESULTS
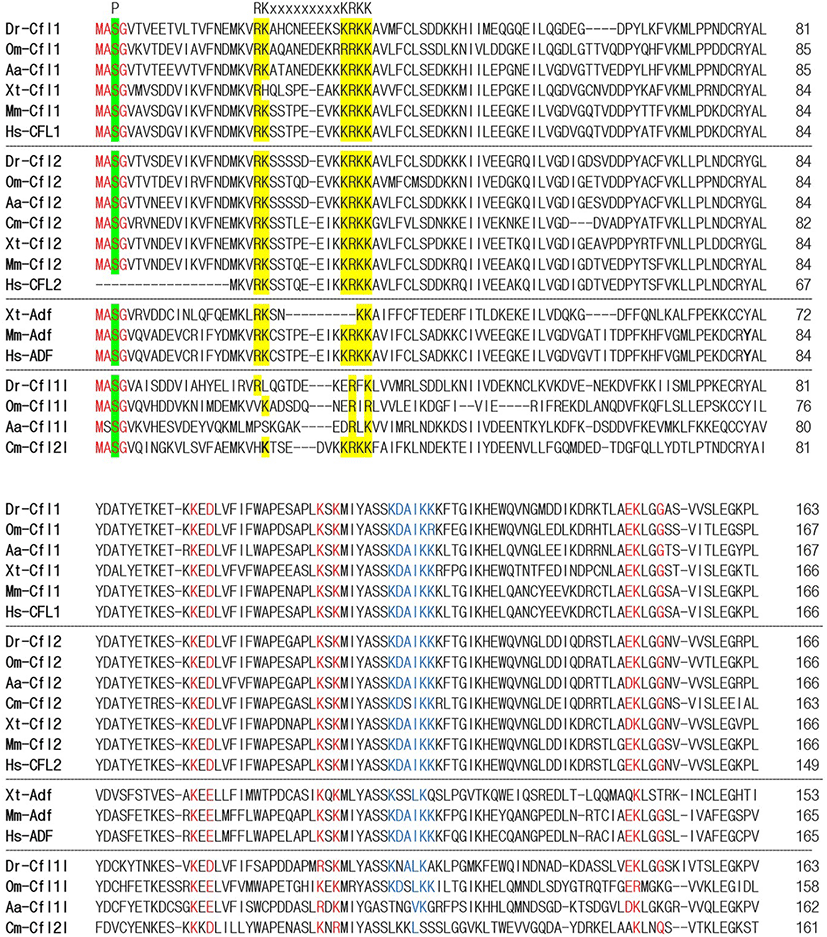
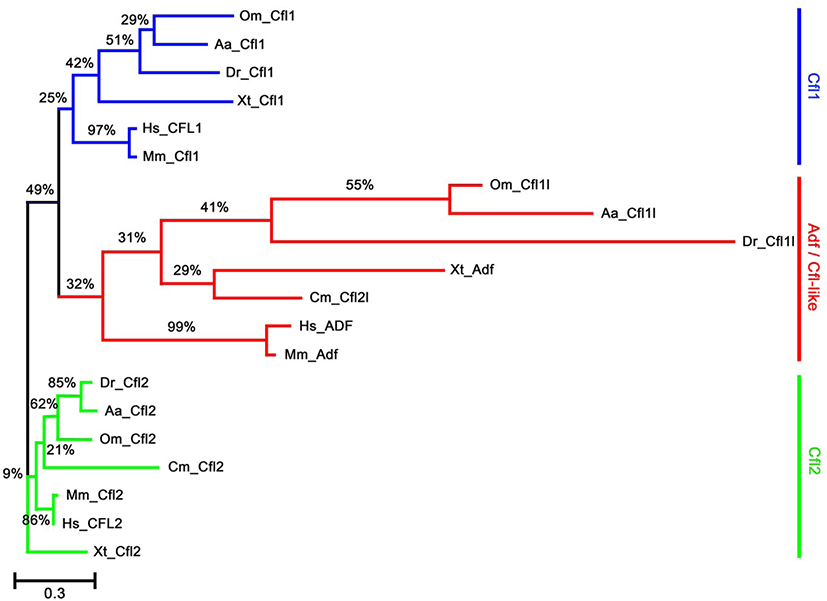
To identify the orthologs of zebrafish cfl1l in vertebrates, we searched the NCBI database. Apparently, cfl1l genes were only found in the fish genomes, including medaka and European eel, with cfl2l genes identified in sharks. Interestingly, orthologs of the adf gene were not identified in the fish genomes bearing the cfl1l or cfl2l gene. Compared to the vertebrate cfl and adf genes, fish cfl1l and cfl2l genes shared most residues essential for interactions of yeast Cfl with actin (red in Fig. 1; Lappalainen et al., 1997) as well as a phosphorylation site at Serine-3 (highlighted in green in Fig. 1; Agnew et al., 1995; Moriyama et al., 1996). Still, they showed some variability in the region that was also important for actin interactions (blue in Fig. 1; Yonezawa et al., 1989) as well as in the nuclear localization signal (NLS) (highlighted in yellow in Fig. 1; Munsie et al., 2012). To verify vertebrate orthologs of zebrafish cfl1l, we carried out the phylogenetic analysis with the multiple sequence alignment for amino acids deduced from the CDS of vertebrate cfl and adf genes (Fig. 1). Interestingly, fish cfl-like (cfl1l and cfl2l) genes belonged to the vertebrate adf group with a 32% bootstrap value (red lines in Fig. 2). Fish cfl1 and cfl2 genes were also grouped with vertebrate cfl1 and cfl2 genes, respectively, with lower bootstrap values than the adf group (blue and green lines, respectively in Fig. 2). Although the bootstrap values are low, our preliminary phylogenetic analysis of vertebrate cfl and adf genes suggests that the fish cfl-like genes might be orthologs of the vertebrate adf gene.
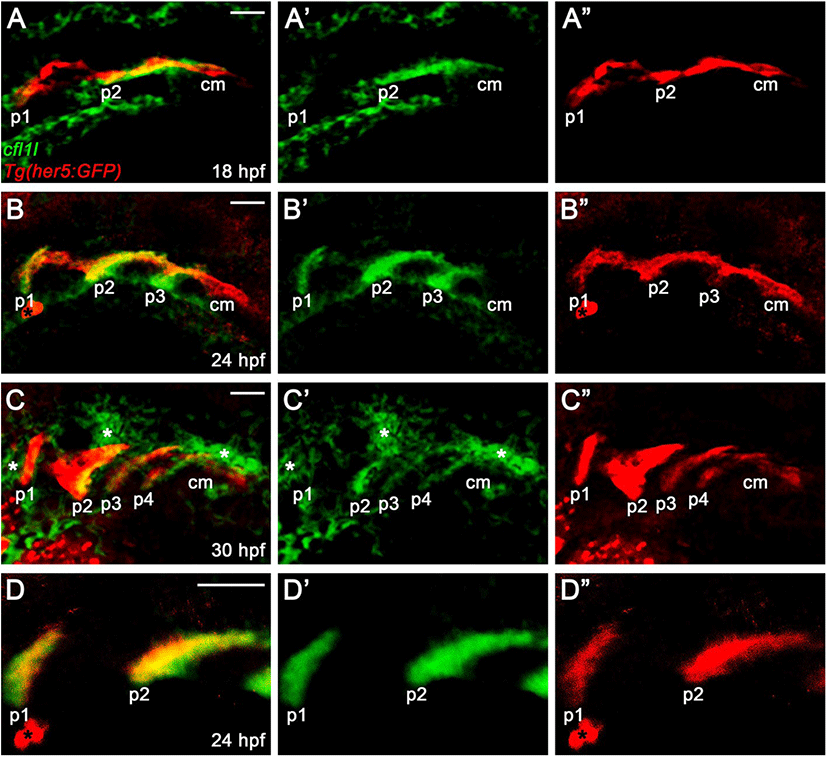
In zebrafish, cfl1 is required to develop the heart and kidney, with cfl2 being implicated in heart development, whereas the physiological role of cfl1l has not been reported yet (Ashworth et al., 2010; Fukuda et al., 2019). Recently, a single-cell RNA sequencing in zebrafish revealed cfl1l expression in 24 hours-post-fertilization (hpf) old PE cells, implying a potential role of Cfl1l in craniofacial development (Wagner et al., 2018). To investigate the developmental role of cfl1l in zebrafish, we first analyzed the expression of cfl1l during the morphogenesis of pouches. To do so, we performed in situ hybridization for cfl1l in wild-type TU embryos carrying Tg(her5:GFP) transgene, a reporter of the PE and pouches (Tallafuß & Bally-Cuif, 2003). When the first two pouches formed at 18 hpf, cfl1l was expressed in the pouches, with its expression in the second pouch being intense (Fig. 3A). However, cfl1l expression in the posterior cell mass was not seen, in which future pouches formed (Fig. 3A). As the third pouch developed at 24 hpf, cfl11 expression in the pouches was evident, whereas the posterior cell mass still did not express cfl1l (Fig. 3B). At 30 hpf, when posterior pouches, including the fourth pouch, were grown out, cfl1l was expressed in the pouches, with the cfl1l expression in the first pouch being abolished. Still, cfl1l was not expressed in the posterior cell mass (Fig. 3C). Besides, cfl1l expression was observed in other tissues adjacent to the pouches at 30 hpf (asterisks in Fig. 3C). We verified cfl1l expression in the pouches with a high magnification single section image showing colocalization of cfl1l transcripts with GFP expressed in her5-positive pouches (Fig. 3D). In summary, cfl1l was expressed sequentially in the pouches during pouch formation but was not expressed in the posterior cell mass, in which pouch-forming cells had not formed pouches yet.
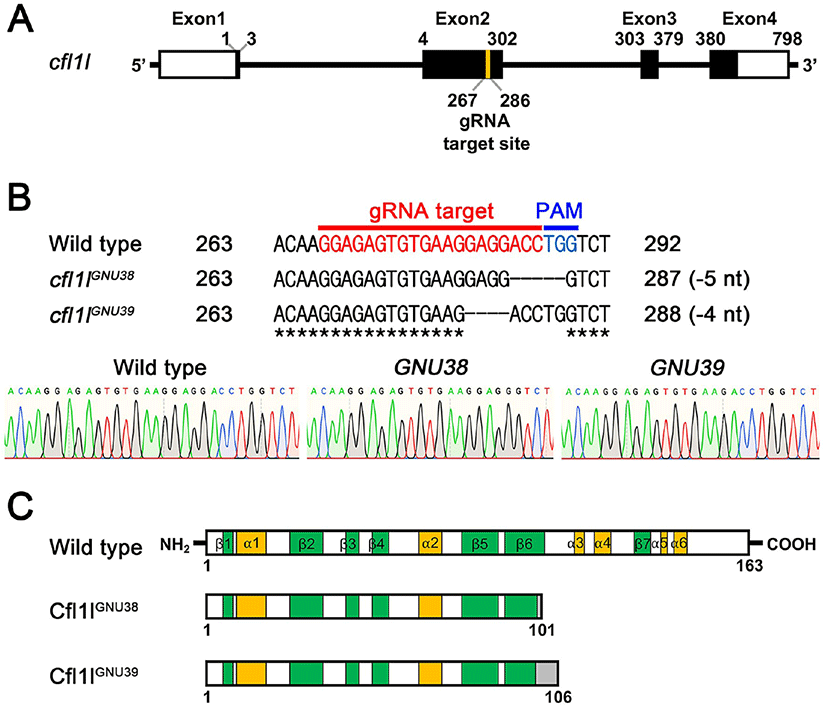
Given the essential role of pharyngeal pouches in craniofacial development (Piotrowski & Nüsslein-Volhard, 2000; Lindsay et al., 2001; Tran et al., 2011), cfl1l expression in the pouches could be necessary to develop pouch itself and/or facial skeletons. To analyze the function of cfl1l in craniofacial development, we generated loss-of-function mutations in the cfl1l gene with CRISPR/ Cas9 system. We designed a gRNA targeting the second exon among four exons of the cfl1l gene with ZiFIT (Fig. 4A; Baker, 2014). We established two mutant cfl1lGNU38 and cfl1lGNU39 alleles in which five and four nucleotides were deleted in the target region, respectively (Fig. 4B). While the wild-type cfl1l gene was expected to encode 163 amino acids bearing six α-helixes and seven β-sheets in the secondary structure of Cfl1l, both cfl1lGNU38 and cfl1lGNU39 alleles were predicted to encode 101 and 106 amino acids, respectively, with the resulting mutant Cfl1lGNU38 and Cfl1lGNU39 proteins being truncated at the sixth β-sheet due to the misframed amino acids and an early termination codon (Fig. 4C). Considering the importance of the sixth β-sheet and the following two α-helixes as a central part in the ADF-H domain of ADF/Cofilin superfamily proteins (Lappalainen et al., 1998; Ono, 2007), Cfl1lGNU38 and Cfl1lGNU39 mutant proteins lacking the sixth β-sheet and the following two α-helixes, were expected to be less functional than wild-type Cfl1l protein. Thus, we suggest that cfl1lGNU38 and cfl1lGNU39 are hypomorphic alleles.
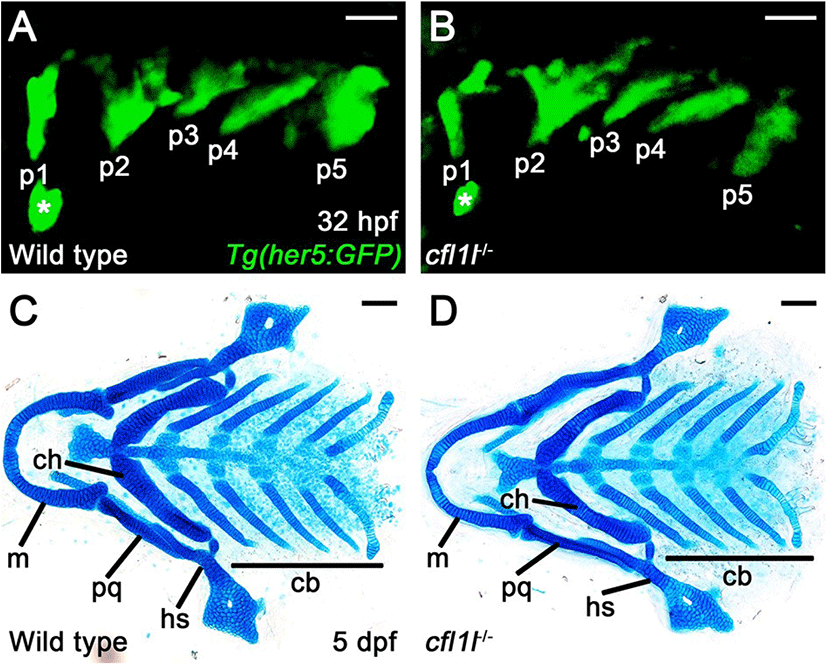
To access the function of cfl1l in craniofacial development, we first examined the pouches in single mutants of cfl1lGNU38 and cfl1lGNU39 harboring Tg(her5:GFP) transgene that allowed us to visualize the pouches during the morphogenesis of pouches directly. In wild-type siblings, pouch formation was completed at 32 hpf, with five pouches seen by the Tg(her5:GFP) transgene (Fig. 5A). Like wild types, five pouches were observed in cfl1l mutants by the Tg(her5:GFP) transgene (Fig. 5B). We have analyzed pouch formation in 64 cfl1l mutant animals, with none showing defects in pouches at 32 hpf, which suggests that cfl1l is not essential for pouch development. Since genes expressed in the pouches could be required for facial skeleton development rather than pouch formation (Miller et al., 2000; Zuniga et al., 2010), we next analyzed facial cartilages in cfl1l mutants at 5 dpf. We examined facial cartilages in 286 wild-type siblings and 93 cfl1l mutants. In wild-type siblings and cfl1l mutants, all elements of facial cartilages driven from the pharyngeal arches were normal, including the hyosymplectic (hs) and ceratobranchial (cb) cartilages whose formation was dependent upon appropriate pouch development (Fig. 5C,D). Even though cfl1lGNU38 and cfl1lGNU39 are likely hypomorphic rather than null alleles, normal development of the pouches and facial cartilages seen in cfl1l mutants suggests that cfl1l expression in the pouches might not be essential for craniofacial development, such as the pouches and facial skeletons, in zebrafish.
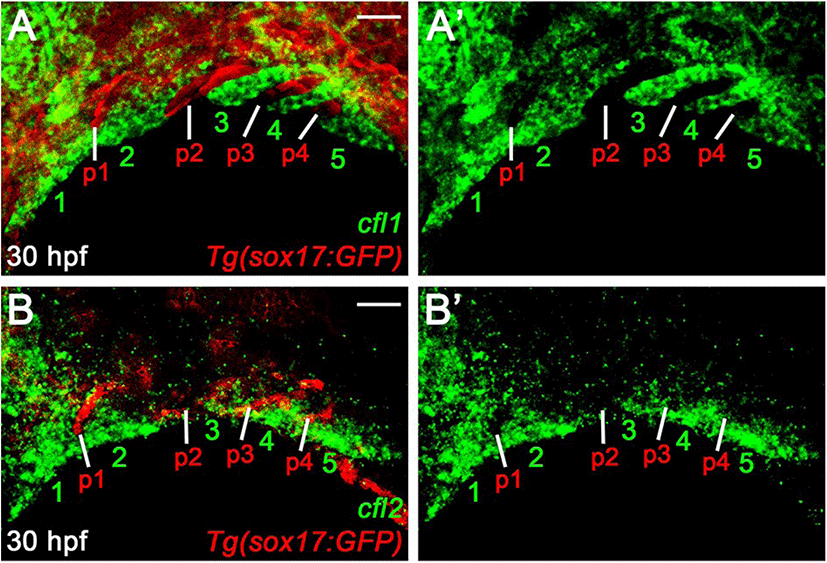
The normal pouches and facial cartilages observed in cfl1l mutants could be a consequence of the genetic redundancy of Cfl1l with Cfl1 or Cfl2 for craniofacial development. If this is the case, cfl1l expression would overlap with cfl1 or cfl2 in the pouches during craniofacial development. To examine a potential redundancy among cfl genes for pouch formation, we analyzed the expression of cfl1 and cfl2 in the pharyngeal regions. At 30 hpf, cfl1 was expressed in the pharyngeal arches rather than sox17-positive pouches (numbers in Fig. 6A). Similar to cfl1, cfl2 was expressed in the ventral domains of the arches, with its expression in sox17-positive pouches being barely seen (numbers in Fig. 6B). Expression of cfl1 and cfl2 in the arches adjacent to the pouches at 30 hpf, suggests that Cfl1l is unlikely to be redundant with Cfl1 or Cfl2 for pouch formation.
DISCUSSION
In this study, we analyzed the expression and function of cfl1l, which might be an ortholog of vertebrate adf, in craniofacial development in zebrafish. While cfl1l was expressed in the pouches during the morphogenesis of pouches, loss-of-function mutations in cfl1l did not affect the development of pouches and facial cartilages. Given the importance of ADF/Cfl family proteins in actin cytoskeleton dynamics, together with the essential role of actin cytoskeleton dynamics in pouch formation (Quinlan et al., 2004; Ono, 2007; Shishkin et al., 2016), the normal pouches seen in cfl1l mutants were unexpected. The normal pouches could be due to the genetic redundancy of Cfl1l with Cfl1 or Cfl2. Indeed, in mice, Adf and Cfl1 show redundant requirements for kidney development, with kidney-specific Cfl1 knockout mice showing normal kidney (Kuure et al., 2010). Although the function of Cfl1 and Cfl2 in heart development remains undetermined, Cfl1 and Cfl2 are expressed together in mammalian cardiomyocytes, implying a possibility of genetic redundancy of Cfl1 and Cfl2 in heart development (Kremneva et al., 2014). However, our analysis of the expression of cfl1 and cfl2 during craniofacial development suggests that the genetic redundancy among Cfls is unlikely in pouch formation. The normal pouches observed in cfl1l mutant fish might be due to the independence of actin cytoskeleton dynamics from Cfl1l in pouch-forming cells. In vertebrate cells, actin cytoskeleton dynamics are achieved by not only ADF/Cfl family proteins but also other actin regulators, including the actin regulator Actin related protein 2/3 (Arp2/3) complex and neural Wiskott-Aldrich syndrome protein (N-WASP, currently renamed as WASP like actin nucleation promoting factor [WASL]). In Xenopus extracts, Cdc42 regulates actin cytoskeleton dynamics through the Arp2/3 complex activated by WASL (Rohatgi et al., 1999). In zebrafish, the Arp2/3 is necessary for proper lamellipodia-like protrusion formation in the migrating posterior lateral primordium through actin dynamics (Olson & Nechiporuk, 2021). Moreover, Wasl acting downstream of Cdc42 is involved in the assembly of endothelial filopodia through actin remodeling in zebrafish (Wakayama et al., 2015). In zebrafish pouch formation, Cdc42 is required to rearrange pouch-forming cells (Choe et al., 2013). Interestingly, in the Zebrafish Model Organism Database (ZFIN), we have found that actin related protein 2/3 complex (arpc) genes composing the Arp2/3 complex are expressed ubiquitously in most embryonic tissues or the pharyngeal regions, with wasl-b being expressed in the pharyngeal regions, during pouch formation. Determination of the specific expression domains of the arpc and wasl-b genes in the pharyngeal regions, followed by the genetic analysis of these genes with cfl1l in pouch formation, will provide a better insight into the function of cfl1l in craniofacial development.