INTRODUCTION
Spermatogenesis is a complicated and particularized process that creates haploid spermatid and sperm from initially undifferentiated germ cells such as spermatogonia (França et al., 1999; Calvo et al., 2000; Pinart et al., 2000; L’Hernault, 2006). In mammal, the spermatogenesis has been known to be regulated by intrinsic gene expression and extrinsic stimuli as hormonal variation and temperature. The seminiferous tubule is specific tubular structures in the testis and it is developed from testis cord of embryonic gonads (Jost et al., 1981). At birth, the seminiferous epithelia are consisted of Sertoli cells and gonocytes only, and the first stage of spermatogenesis initiates a few days after birth. The spermatogenesis is completed around PD 37 in mice with the forming sperm (Clermont & Perey, 1957; Drumond et al., 2011), and it is divided into 12 phased based on the associations of germ cells in four stages of spermatogenesis (Clermont & Leblond, 1953; Ahmed & de Rooij, 2009; Nakata et al., 2015). The first stage (spermatocytogenesis; mitosis) of spermatogenesis is forming spermatogonial stem cells as spermatogonium, and it highly expressed PLZF, SALL4, and CDH1 (Costoya et al., 2004; Tokuda et al., 2007; Gassei & Orwig, 2013). The PLZF is the first transcription factor to be discovered as being involved in spermatogonial self-renewal. The second stage (spermatidogenesis; meiosis I, II) is forming spermatocyte which undergoes meiosis, a production of haploid gametes and maintaining genome integrity. The SCP3 is a marker and essential for synaptonemal complex formation during meiosis in spermatocyte (Yuan et al., 2000). The third stage (spermiogenesis) is forming spermatids which marker is PNA, and final stage (spermiation) is forming spermatozoids (sperm) (L’Hernault, 2006). All stages of spermatogenesis are closely regulated by Sertoli cells (Griswold, 1998). Although many studies reported the signal pathway of spermatogenesis, there is a need to study the mechanism of gene regulation of spermatogenesis.
Cyclic-Amp response element binding protein zhangfei (CREBZF) is a novel regulator of various cellular mechanisms such as unfolded protein response, apoptosis, cell growth, migration, and differentiation by direct interaction to transcription factors due to lack of DNA binding domain (Zhang et al., 2010; Jang et al., 2014; Zhang & Misra, 2014; Jang et al., 2015; Zhang et al., 2015). Also, recent study reported that CREBZF is regulated by sexual hormones in mouse uterus (Lin et al., 2013). However, the study of CREBZF in spermatogenesis of testis is not discovered, yet.
In this study, to identify the function of CREBZF, the expression level of CREBZF in mouse tissues was examined. Also, the specific localization of CREBZF in seminiferous tubule of testis was firstly presented.
MATERIALS AND METHODS
All animal experiments were approved by Institutional Agricultural Animal Care and Use Committee of CHA University (IACUC No. 160018) and conducted according to the guidelines for the care and use of laboratory animals. The animal studies were performed with 3 of PD 7, 10, 14, 21, and 42 day male ICR mice from KOATECH (Korea). Mice were housed in animal care facility at the CHA University of CHA biocomplex with a temperature-controlled environment under 12-hours dark-light cycle and free access to standard rodent diet and water.
To extract total RNA from each tissues and testes, mice were sacrificed and isolated respectively. The tissues were placed into a vessel and homogenized in Trizol reagent (Invitrogen, USA) using homogenizer. The total RNA extraction was according to manufacturer’s protocol. cDNA synthesis was performed using SensiFAST cDNA Synthesis Kit (Bioline, UK) according to manufacturer’s protocol, and the thermal cycling conditions were as follows: reverse transcription at 45℃ for 60 minutes, and 4℃ for 10 minutes. Real-time PCR and RT-PCR analyses were performed in triplicate using Solg Taq DNA polymerase (Solgent, Korea) and iQ SYBR Green Supermix (Bio-Rad, USA). The thermal cycling conditions were as follows: initial denaturation at 95℃ for 3 minutes, followed by 40 cycles of 95℃ for 10 seconds, 60℃ for 15 seconds, and 72℃ for 20 seconds. Expression levels were normalized to those of endogenous GAPDH. The primer sequences were as follows: mouse CREBZF forward 3’-CTGCCCGTCT TAATC GGCTC-5ʼ; reverse 3ʼ-CCGTAGGTAGCGACTCTCCTC-5ʼ, mouse GAPDH forward 3’-AGGTCGGTGTGAACGGAT TTG-5ʼ; reverse 3ʼ-TGTAGACCATGTAGTTGA GGT- 5ʼ.
Testes were fixed for 1 week in 4 % formaldehyde at 4℃ and embedded in paraffin. The paraffin blocks were sectioned at 5 μm of thickness using a microtome and put on microscope slides (HistoBond, Germany). For immunostaining, sections were deparaffinized as follows; slides were dipped three times for 5 minutes in xylene (Biosesang, Korea), two times for 5 minutes in 95 % ethanol, one time for 5 minutes in 90 %, 80 %, 70 %, 50% ethanol, and 10 minutes in distilled water. For immunofluorescence, deparaffinized slides were placed into an antigen Retrieval solution (IHCworld, USA), and antigen retrieval was performed using a Retrieval steamer (IHCworld, USA) according to the manufacturer’s protocol. After washing in PBS, excess PBS was removed, and blocking buffer (4% BSA and 5% rabbit serum in PBS) was added to the slides. The slides were incubated in a humidified chamber for 4 hours at room temperature (RT). Then, the slides were incubated with the following primary antibodies for 16 hours at 4℃: Goat polyclonal antibody against CREBZF (1: 1000, Santacruz), mouse monoclonal anti-DEAD-Box Helicase 4 (DDX4, 1:500, Abcam), mouse monoclonal anti-Promyelocytic leukemia zinc finger protein (PLZF, 1:500, Santacruz), mouse monoclonal anti-Synaptonemal complex Protein 3 (SCP3, 1:200, Abcam), rabbit polyclonal anti-GATA binding protein 4 (GATA4, 1:300, Abcam). After three washes in PBS, the slides were incubated with Alexa 488 mouse anti-Goat and Alexa 546 mouse anti-rabbit (1:1500, Invitrogen, UK) for 2 hours at RT. A 4’,6-diamidino-2-phenylindole (DAPI, 1:20000, Life Technologies, USA) was used to stain the nuclei. Mounting medium (DAKO, USA) was applied to the tissue sections prior to covering them with glass coverslips. All images were obtained using a confocal microscope (Leica) and analyzed by the imaging software LAS lite (Leica).
RESULTS
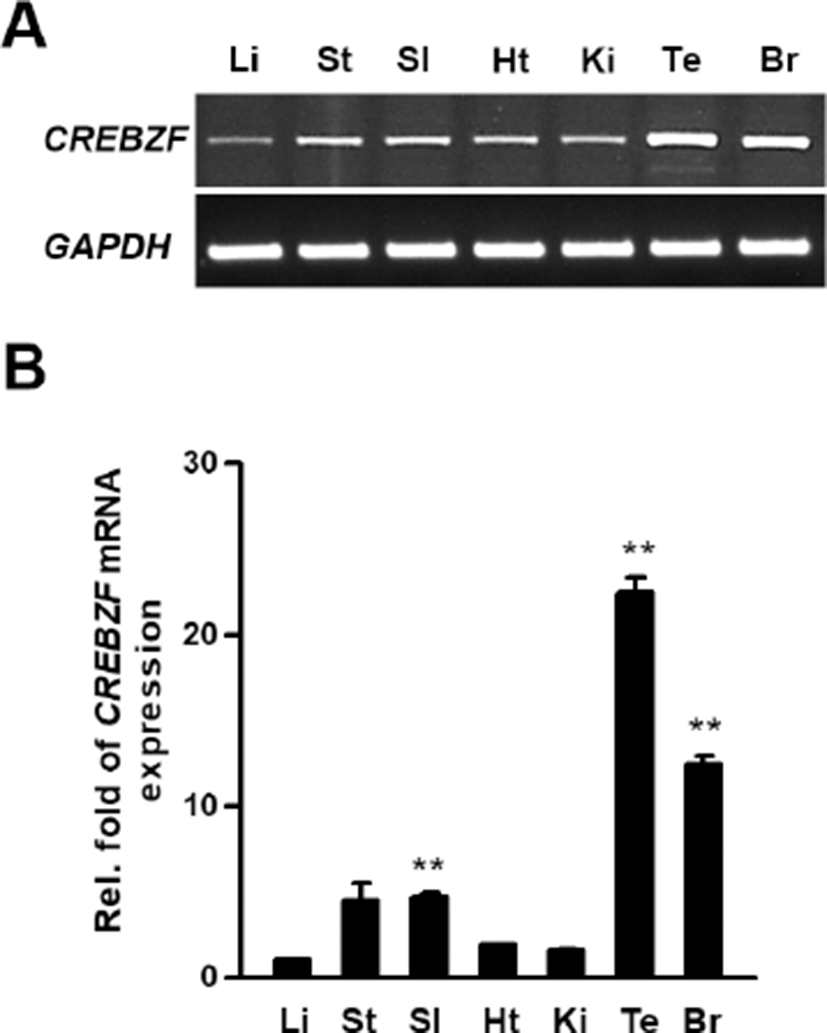
To examine expression levels of CREBZF mRNA in various tissues, we conducted RT-PCR and qRT-PCR using respective cDNA from 6-weeks old mice organs with designed mouse CREBZF primers as described. The CREBZF transcript was highly expressed in testis than other tissues in adult mice (Fig. 1A and 1B). These results showed that CREBZF may have an important role in testis for reproduction.
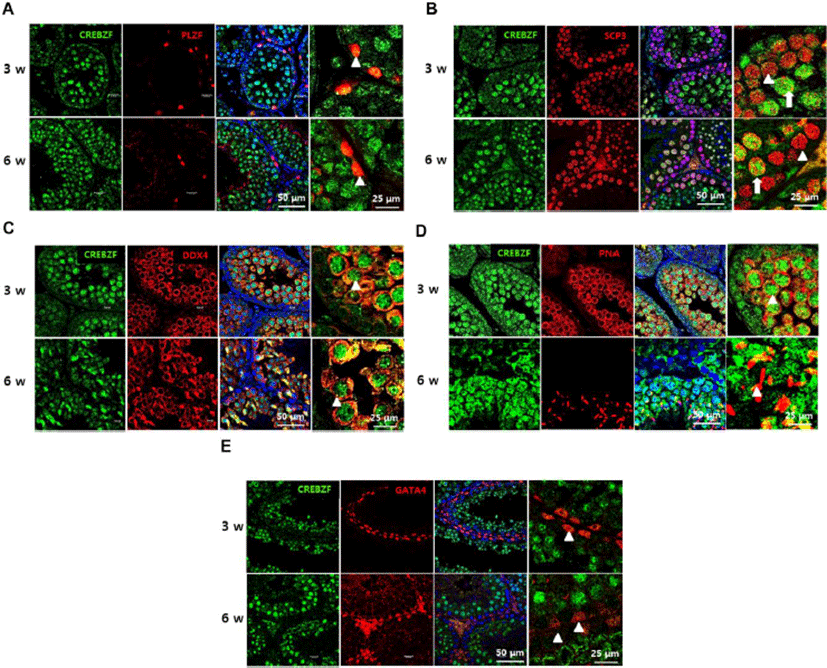
To evaluate expression patterns of CREBZF in the seminiferous tubule, immunofluorescence was performed using anti-CREBZF with anti-PLZF, a marker of spermatogonia, anti-SCP3, a marker of spermatocyte, anti-DDX4, a marker of germ cells, anti-PNA, a marker of spermatid, and anti-GATA4, a marker of sertoli cells for double staining. The results showed that CREBZF was not expressed on spermatogonia (Fig. 2A), whereas highly expressed on meiotic reproductive cells (Fig. 2C), such as spermatocyte (Fig. 2B) and spermatid (Fig. 2D). Taken together, these results indicate that CREBZF is specifically expressed on the germ cells progressing to meiosis, but less expression in spermatogonia and Sertoli cells.
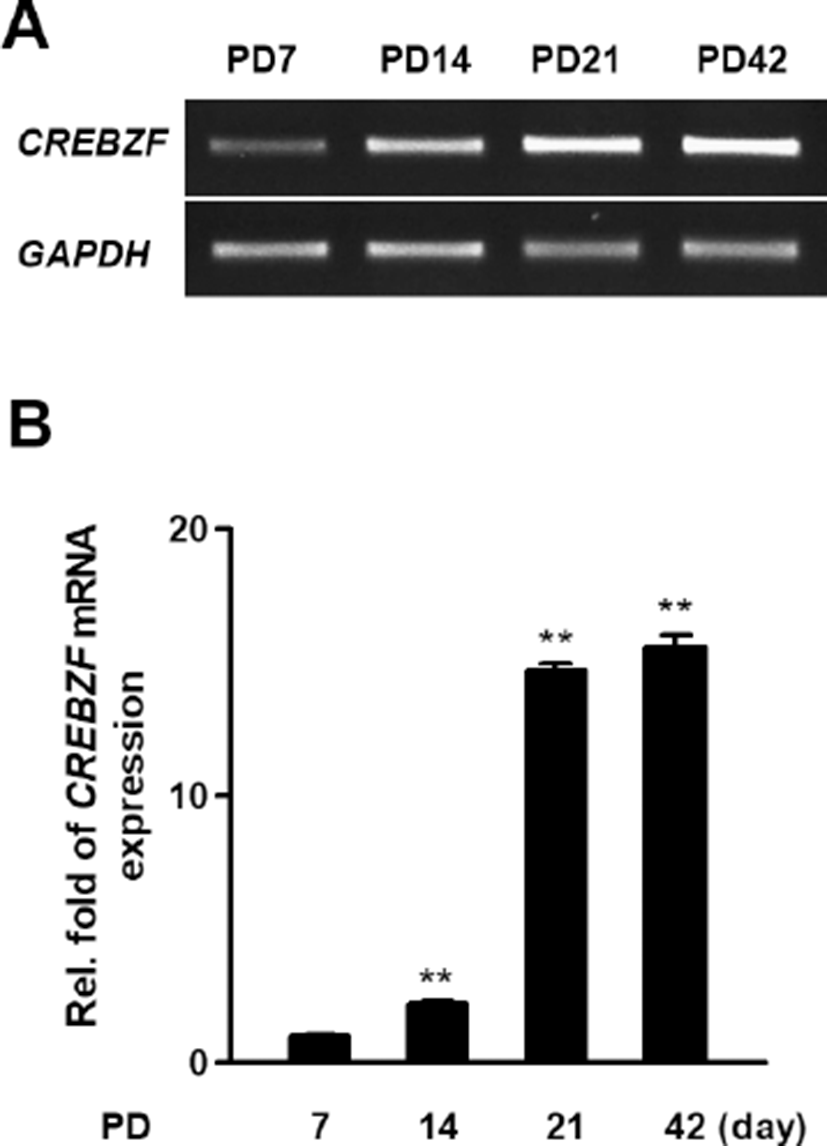
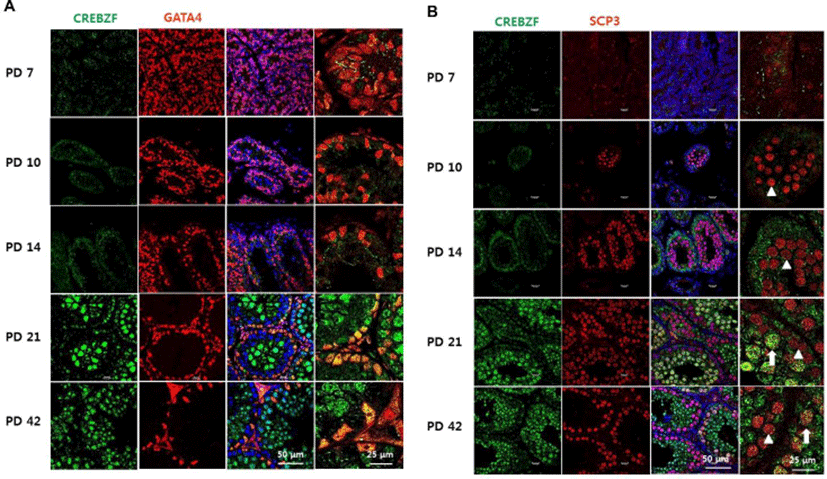
To investigate whether CREBZF is changed during development of testis, we examined the mRNA expression of CREBZF in PD 7, PD 14, PD 21, and PD 42 mouse testis. RT-PCR and qRT-PCR showed CREBZF transcript gradually elevated during postnatal development. Especially, that result indicated that the expression of CREBZF transcript dramatically increased from PD 21 (Fig. 3A, B). To further confirm the location of CREBZF in seminiferous tubule, double staining of immunofluorescence was performed using GATA4 and SCP3. The confocal imaging analysis identified that CREBZF protein was no expressed until PD 14, and dramatically increased from PD 21. However, GATA4 was highly expressed on the whole cells of testes in PD 7 and gradually decreased during spermatogenesis (Fig. 4A). Expression of SCP3 was initiated in PD 10 and highly expressed in whole meiotic cells, whereas expression of CREBZF was initiated in PD 21 (Fig. 4B). These results indicated that CREBZF expression is closely related to meiosis during spermatogenesis.
DISCUSSION
In this study, CREBZF was highly expressed in testis and specifically located on germ cells of seminiferous tubule in mouse testis. There was no expression of CREBZF in new born mouse testis. During postnatal development, the expression of CREBZF increased significantly. In adult mouse, expression of CREBZF was low in spermatogonia and Sertoli cells but strong in spermatocyte and spermatid. Especially, CREBZF showed strong expression only in certain cells of spermatocytes. Therefore, it is expected that CREBZF may play an important role in spermatogenesis.
The spermatogenesis differs depending on the type of cells in the seminiferous tubule between hibernating or non-hibernating mammals. (Barnes et al., 1986; Jagiello et al., 1986; Parua et al., 2011; Lee, 2013). In mouse, seminiferous epithelium is divided into spermatogonium, spermatocytes, spermatid, sperm, and Sertoli cells. There was a need to determine whether CREBZF is mainly expressed in certain cells of the seminiferous epithelium because the CREBZF has various function during cell metabolism and development in somatic cells such as hepatocyte, osteoblast, adipocyte, and some of cancer cells (Zhang & Misra, 2014; Jang et al., 2014; Jang et al., 2015). In this study showed the specific expression of CREBZF in cells which are presumed to be secondary spermatocytes (Fig. 2B). There are two types of spermatocyte as primary and secondary spermatocytes and are differentiated from spermatogonia via process of spermatogenesis. Primary spermatocytes are diploid (2N) which are located on cortex of tubules near spermatogonia, and formed from spermatogonia (Griswold, 1998; L’Hernault, 2006). After meiosis I, two secondary spermatocytes (haploid, N) are formed and undergo meiosis II which is located near spermatids (Pinart et al., 2000; L’Hernault, 2006). Previous study reported that the meiosis is started about PD 10 in mouse testis (Yuan et al., 2000; L’Hernault, 2006). In our results, the SCP3, a marker of spermatocyte, expression is initiated at PD 10, but CREBZF is initiated at PD 21 (Fig. 4B). Based on the fact, it can be inferred that CREBZF may play an important role in the late meiosis stage (meiosis II), not in the early meiosis stage (meiosis I).
The Sertoli cells provide important factors for successful progression of spermatogenesis (Griswold, 1998). However, CREBZF was not detected on Sertoli cells in our results. Since CREBZF has been reported to play an important role in various cells, it was necessary to further confirmed whether CREBZF has not working in Sertoli cells during postnatal development. At PD 7, seminiferous epithelial cells are composed of most Sertoli cells, which are GATA4-positive somatic cells, and a few gonocytes in mouse testis (Griswold, 1998). The CREBZF protein expression was not detected on this stage of PD 7, and during postnatal development, GATA4-positive cells were not matched with CREBZF-positive cells. These results suggest that CREBZF play critical roles in only germ cells during meiosis.
In conclusion, CREBZF is highly expressed in adult mouse testis than other tissues, and it is specifically expressed on later stage of spermatocyte in seminiferous tubule of testis. In addition, the CREBZF is not expressed on spermatogonia and Sertoli cells. Further study will be performed to determine detailed functions of CREBZF during meiosis of spermatogenesis.