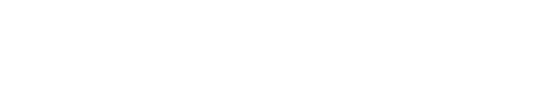INTRODUCTION
Reproduction is a strategy used by individuals to maintain a species, thus a population. Individuals adjust their behavior to the ever-changing ambient environment, in order to secure the survival of their own as well as of the descendants. For example, different species have developed various reproductive strategies to account for seasonal changes (Choi & Lee, 2012; Dardente et al., 2016). As the environment changes throughout evolution, the living patterns change accordingly. The diversification and stabilization of living patterns over generations can lead to the advent of new species. In fact, it is speculated that all extant species underwent such process to arrive at their current state. Since the real entity that gets transmitted to the future generations is genes, it is possible that the genes of the common ancestors were specialized to allow for the formation of diversity. Thus, speciation, which forms numerous new species, is an ongoing evolutionary phenomenon.
It is well documented that small modifications of the inheritable traits have always existed and conserved over generations. These modifications are made on the genes, by genetic recombinations during prophase of meiosis for the formation of germ cells and development of mutations during duplication. Altered genetic traits then lead to differentiations and diversity among species. These steps compose the speciation process observed in living organisms.
Another factor causing evolution is the applicability of the inheritable traits to the encircling environment. It has long been established that in order for a species to survive during the adaptation process, it is necessary for favorable traits to be transmitted to next generations whereas unfavorable traits disappear. As the environment varies depending on the geographic location, natural selection exerts different effects on the likelihood of favorable traits to be passed down. If inheritable traits do not adapt properly, abrupt changes in the environment could drive the population to extinction.
Evolutionary biology originates from the concept of natural selection proposed by Charles Darwin and other scientists. The theory was widely known by the book titled “The Origin of Species” published in 1859 by Charles Darwin (Darwin, 1859). He observed and described in great detail the organisms and fossils inhabiting the regions he visited via the “Beagle,” as well as the geological characteristics (Darwin, 1839; Darwin, 1891). It is possible that the finches in the Galapagos Islands provided Darwin with an idea about evolution, although he had not fully formulated the concept at that time. He eventually came to realize that, the obvious variations in the body size, beak size and shape of the finches arise from variations in the geographical environment, or ecological niche. The beaks with favorable traits were naturally selected for generation after generation, until they ramified out to form new species (Skinner et al., 2014; McNew et al., 2017). Unlike other birds that were almost identical to mainland finches in South America in every aspect, Darwin's finches displayed adaptations to the type of food they ate in order to accommodate different niches on the Galapagos Islands. In this case, the adaptive trait that got transmitted to the offspring provides benefits.
In cells, gene expression can be altered without having the sequence of nucleic acid itself modified. Known as ‘Epigenetics,’ the most common form of histone modification is methylation, which has been poorly understood (Felsenfeld, 2014). Regardless of the sequence, the epigenetic modifications regulate the genome and are stable during mitosis. Some modifications are stable during meiosis, which allows them to be transmitted through the germ cell lineage and be inserted into the chromosomes (Jirtle & Skinner, 2007). DNA methylation as an epigenetic mechanism can be imprinted and inherited from one generation to another, thus having an evolutionary impact. Similarly to genetic changes containing recombinant genes in the meiotic division and mutations, epigenetic modifications can play an important role in evolutionary processes (Guerrero-Bosagna et al., 2010; Ben Maamar et al., 2018) - Darwin's finches are an example of this.
The reproductive activity essential to sustaining a species requires a great amount of energy, which is a course not to be avoidable. The hypothalamus-pituitary-gonadal axis in vertebrates is the master regulator of the reproductive endocrine system. Gonadotropin-releasing hormone (GnRH), which is synthesized in the hypothalamus and released into hypothalamo-pituitary portal blood vessel, acts on the cells of the anterior pituitary to induce the secretion of follicle stimulating hormone (FSH) and luteinizing hormone (LH) simultaneously. These two gonadotropins operate on the male gonad testis to promote the formation of spermatozoa and to synthesize male sex steroid hormone, testosterone. In females, they act on female go-nad ovary to foster the maturation of the ovum and simultaneously to produce female sex hormone estrogen (Charlton et al., 1983; Mason et al., 1986; Schwanzel-Fukuda et al., 1989). Both sex hormones then regulate the release of GnRH from hypothalamus via negative feedback. The sperm and ovum combine to form the offspring and the genes involved are transmitted to the next generations. The most important substance in this entire process is the GnRH.
In general, a hypothalamus-derived neural peptide cau-ses the pituitary to release its corresponding hormone. The case of the hypothalamic hormone GnRH is peculiar, because it causes the pituitary to release two hormones - FSH and LH. Moreover, it was discovered that activin and inhibin found in the gonad regulate the release of FSH (Carroll et al., 1989; Weiss et al., 1993). A specific substance to control LH release was not found at the time. Interestingly, a hypothalamic factor that inhibits the secretion of GnRH was discovered in birds and named gonadotropin inhibiting hormone (GnIH) (Tsutsui et al., 2000). Since then, GnIH has been identified in all vertebrates, including humans (Tsutsui et al., 2017). Thus, the mechanism by which the reproductive endocrine system is regulated was realized to be far more complicated.
The present research analyzes how GnRH, the most basic yet important regulatory substance of reproductive endocrine system, has transformed through evolution of Chordata. Changes in GnRH amino acid sequences are discussed first, then changes in GnRH nucleic acid sequences. These analyses reveal the evolutionary viewpoint on GnRH of Chordata.
DISCOVERY AND VARIATIONS
GnRH was first isolated from the hypothalamus of mammals in early 1970s (Baba et al., 1971; Matsuo et al., 1971; Schally et al., 1971). It was then identified in most animals of Chordata and subsequently in invertebrates (Millar, 2005; Roch et al., 2011; Hasunuma & Terakado, 2013; Sakai et al., 2017). The functional hypothalamic GnRH found in vertebrates is a decapeptide, consisting of 10 amino acids (Fig. 1). The N-terminal has pyroglutamate (pGlu), which has a ring shape and is formed by modified glutamine. The C-terminal containing an amine group is composed of carboxamide. For example, the amino acid sequence of human GnRH is pGlu-His-Trp-Ser-Tyr-Gly-Leu-Arg-Pro-Gly-NH2 (QHWSYGLRPG in one letter code). The original GnRH discovered was found to be related to reproduction and named GnRH1. Following GnRH1, other forms of GnRH were continuously uncovered (Table 1).
| Order | GnRH1 | GnRH2 | GnRH3 | Sources |
|---|---|---|---|---|
| Primates | QHWSYGLRPG | ② | NG_016457.1, AF036329.1 | |
| Scandentia | QHWSYGLRPG | ② | U63326.1, U63327.1 | |
| Rodentia | QHWSYGLRPG | U91938.1 | ||
| QHWSFGLSPG | AC105786.3 | |||
| QYWSYGVRPG | NM_001172956.1 | |||
| Soricomorpha | ② | AF107315.1 | ||
| Diprotodontia | QHWSYGLRPG | ② | KJ191755.1, KJ191756.1 | |
| Passeriformes | QHWSYGLQPG | JN797488.1 | ||
| Anseriformes | QHWSYGLQPG | DQ023158.1 | ||
| Columbiformes | QHWSYGLQPG | EU699771.1 | ||
| Galliformes | QHWSYGLQPG | ② | JN609567.1, JN675400.1 | |
| Squamata | ② | AB104485.1 | ||
| Testudines | ② | XM_008169613.1 | ||
| Anura | QHWSYGLRPG | ② | L28040.1, AF139911.1 | |
| QHWSYGLWPG | FM162564.1 | |||
| Gymnophiona | ② | AF167558.1 | ||
| Scorpaeniformes | QHWSYGLNPG | ② | ③ | JQ028870.1, JQ664745.1, Q724851.1 |
| Perciformes | QHWSYGLSPG | ② | ③ | AB101665.1, KC896411.1, FJ380047.1 |
| QHWSFGLSPG | KM232949.1 | |||
| QLWSYGLSPG | KP159744.1 | |||
| Tetraodontiformes | QHWSYGLSPG | ② | ③ | AB212811.1, AB212813.1, AB212814.1 |
| Cyprinodontiformes | ② | XM_017304681.1 | ||
| Beloniformes | QHWSFGLSPG | ② | ③ | AB041336.1, AB074500.1, AB074501.1 |
| Atheriniformes | QHWSFGLSPG | ② | ③ | AY320285.1, AY744687.1, AY744688.1 |
| Cichliformes | QHWSYGLSPG | ② | ③ | AF076961.4, L27435.1, AF076963.1 |
| Pleuronectiformes | QHWSYGLSPG | ② | ③ | HQ623431.1, HM131601.1, HQ623432.1 |
| Synbranchiformes | ② | AY858054.1 | ||
| Anabantiformes | QHWSYGLSPG | ② | KF113107.1, KF113108.1 | |
| Gobiiformes | ② | XM_020932006.1 | ||
| Scombriformes | QHWSYGLSPG | ② | ③ | HQ108193.1, HQ108194.1, HQ108195.1 |
| Batrachoidiformes | ③ | S79620.1 | ||
| Salmoniformes | QHWSYGMNPG | ② | ③ | AY245104.2, AB365004.1, X79710.1 |
| Siluriformes | QHWSHGLNPG | ② | X78049.1, X78047.1 | |
| Cypriniformes | QHWSRGLSPG | ② | ③ | KM887435.1, BC162951.1, AB020243.1 |
| Clupeiformes | QHWSHGLSPG | ② | ③ | KU323664.1, KU323665.1, KU323666.1 |
| Osteoglossiformes | ② | ③ | AB047326.1, AB047325.1 | |
| Anguilliformes | QHWSYGLRPG | ② | AB026989.1, AB026990.1 | |
| Acipenseriformes | QHWSYGLRPG | ② | EF534707.3, EF534706.2 | |
| Orectolobiformes | ② | XM_020514705.1 | ||
| Chimaeriformes | ② | EF494735.1 | ||
| Petromyzontiformes | QHWSHDWKPG | AY307173.2 | ||
| QHWSHGWFPG | DQ457017.1 | |||
| QHYSLEWKPG | F144481.1 | |||
| Phlebobranchia | QHWSNWWIPG | AB219239.1 | ||
| Stolidobranchia | QHWSYGFSPG | AB682994.1 | ||
| QHWSYGFLPG | Hasunuma and Terakado, 2013 | |||
| Amphioxiformes | QILCARAFTYTHTW | KF601546.1 |
Each residue is represented by a single alphabet. More ancient Orders are found toward the bottom of the table. The amino acid sequences are at the level of Orders in the classification. The sequence of GnRH2 and GnRH3 are QHWSHGWYPG and QHWSYGWLPG, respectively, and are marked accordingly with ② or ③. Sources: GenBank, NCBI Reference Sequence (underlined), and articles cited above. GnRH, gonadotropin-releasing hormone.
As Table 1 indicates, the next form of GnRH was found in chickens and named GnRH2 (Miyamoto et al., 1984; Desaulniers et al., 2017). GnRH2 has the following sequence: pGlu-His-Trp-Ser-His-Gly-Trp-Tyr-Pro-Gly-NH2 (QHWSHGWYPG). This form also induces the secretion of FSH and LH. GnRH2 is well conserved in Chondrichthyes, Actinopterygii, Amphibia, and Reptilia without any modifications in the sequence (Table 1). It is well conserved in Aves and Mammalia as well, although not all have it.
GnRH3 was discovered in lamprey (Table 1). The amino acid sequence of GnRH3 is pGlu-His-Trp-Ser-His-Asp-Trp-Lys-Pro-Gly-NH2 (QHWSYGWLPG) (Sower et al., 1993; Chen & Fernald, 2008; Karigo & Oka, 2013). This GnRH3 is also generally well conserved across different species, except it is only found in a part of Actinopterygii.
Surprisingly, the decapeptide GnRH was detected in the Order Stolidobranchia (Hasunuma & Terakado, 2013). A different form was also reported to be found in Order Phlebobranchia. Both of these displayed similarities to the N- and C-termini of GnRH1, which are the binding sites to the receptor. As for the most primitive Chordata, Leptocardii of Cephalochordata, a possible precursor GnRH-like peptide with similarities to GnRH2 - 14 amino acids and the ability to bind to its receptor - was detected (Roch et al., 2014). This led to the idea that the peptide of Cephalochordata is not associated with GnRH1 that is typically found in Vertebrata.
It has been conjectured that each of the various forms of GnRH can exert its effects on the reproductive endocrine system and has physiological roles. These effects are potentiated upon interaction with their cognate receptors.
GnRH1-releasing neurons are located in the preoptic area of the hypothalamus and principally project onto the anterior pituitary. By controlling the secretion of gonadotropins from the anterior pituitary, GnRH1 plays a major role in regulating reproductive function (Muske et al., 1994; White & Fernald, 1998; Amano et al., 2002). Another role of GnRH1 is to induce the release of growth hormones and somatolactin from the pituitary (Marchant et al., 1989; Kakizawa et al., 1997).
GnRH2 is synthesized in the midbrain tegmentum near the third ventricle. GnRH2 neurons have projections all over the brain, but are especially focused on the midbrain and hindbrain with their axon terminals in the third ventricle (Gonzalez-Martinez et al., 2002; Steven et al., 2003). GnRH1 and GnRH2 can be detected simultaneously in the anterior pituitary, but GnRH1 is thought to be the unique regulator of gonadotropin release (Mongiat et al., 2006).
GnRH3 neurons are only found in Orders of Actinipterygii. They are localized in the terminal nerve ganglion near olfactory bulbs and have projections throughout the whole brain (Chiba et al., 1996; Grens et al., 2005). The function of GnRH3 has been poorly understood, and its roles in reproductive activity and sexual behavior are under examination in various species.
Different forms of GnRH play different roles. GnRH1 is the main peptide regulating the production of germ cell formation and sexual behavior in vertebrates. GnRH2 regulates female reproductive behavior in Actinophetygii, Aves, and Mammalia (Maney et al., 1997; Volkoff & Peter, 1999). GnRH2 is also involved in the control of food intake and energy balance, as well as keeping a balance between survival and reproduction (Temple et al., 2003; Kauffman & Rissman, 2004; Kauffman et al., 2005). Lastly, GnRH3 regulates sexual behaviors in Actinopherygii, such as nest-building, aggression, and spawning (Volkoff & Peter, 1999; Ogawa et al., 2006). It is largely accepted that the primary role of GnRH1 is the direct regulation of reproduction, whereas GnRH2 and GnRH3 modulate behaviors associated with sexual activity.
GnRH AMINO ACID SEQUENCE ANALYSIS
As mentioned above, there are great variations in GnRH1, which governs the overall functioning of reproduction by interacting directly with the reproductive endocrine system (Table 1). In most animals in Chordata, both N- and C-termini of the amino acid sequence of GnRH1 are fairly conserved at positions 1-4, 9, and 10. N-terminal generally consists of pGlu-His-Trp-Ser and C-terminal is fixed to Pro-Gly-NH2. Residues in the middle can vary widely, especially at positions 5, 7, and 8, with the except of the glycine in position 6, which stays constant. As of now, it is unclear why GnRH sequence varies. The N- and C-termini provide structural support for receptor binding and associated biological activity (Millar et al., 2004; Schally et al., 2017). Specifically, pGlu, His, and Trp at N-terminal are essential to physiological activity of GnRH1 and the arginine residue at position 8 strengthens receptor binding (Flanagan & Manilall, 2017). Glycine at position 6 adds flexibility to the sequence, which allows for the two termini to get closer to each other and leads to a more efficient and tighter binding to the receptor (Millar, 2005).
As GnRH gets discovered in more and more animals, there is more information on how amino acids have been modified throughout evolution. The current structure of GnRH1 originates from GnRH-like peptide found in Leptocardii of Cephalochordata (Roch et al., 2014). This peptide was able to bind to GnRH receptors but was 14 amino acids long, which is similar to GnRH2. The decapeptide form of GnRH1 were initially detected in the Order Stolidobranchia, Ascidiacea Tunicata (Hasunuma & Terakado, 2013). Different forms of decapeptide GnRH1 were observed in Phlebobranchia as well. The difference between the two forms found in Stolidobranchia and Phlebobranchia is in the sequence at positions 5-8. Whereas the sequence of the form found in Stolidobranchia goes Tyr-Gly-Phe-Ser or Tyr-Gly-Phe-Leu, the one detected in Phlebobranchia is Arg-Trp-Trp-Ile. Residues at other positions are the same, with Gln-His-Trp-Ser at N-terminal and Pro-Gly at C-terminal.
The residues in the middle of the decapeptide sequence were substituted to a number of different ones in the ancient Order Petromyzontiformes, Actinopterygii of subphylum Vertebrata. Most diverse changes are made on the ones at positions 3 and 5-8. With the emergence of variations in GnRH1 in the Petromyzontiformes, it seems like the residues have undergone a series of trial and error when going to Vertebrata. Presumably, the amino acid sequences that favorably adapted to the environment were chosen by natural selection to be passed down and preserved. In Petromyzontiformes, the 3rd residue was substituted to tyrosine and 5th through 8th residues were altered to Leu-Glu-Trp-Lys, resulting in the sequence QHYSLEWKPG. Other forms observed are QHWSHGWFPG and QHWSHDWKPG. QHW at the N-terminal have come around since early Ascidiacea. Replacing the phenylalanine in QHWSHGWFPG with tyrosine results in GnRH2, whose sequence is QHWSHGWYPG and lineage continues to Primates.
Throughout Actinopterygii, only one or two out of the ten from the sequence were altered and passed down. By the time the decapeptide reached Order Salmoniformes, the 5th residue has changed to tyrosine, which first appeared in primitive Actinopterygii and continued to be the most commonly observed in Vertebrata. In another form of GnRH1, the 7th and 8th residues changed to methionine and asparagine, respectively. No animals before Salmoniformes had the methionine at the 7th position. Consequently, it is speculated that the version containing methionine is associated with the life cycle of salmons, whose environment alternates between fresh water and salt water. In Scombriformes, the 8th residue was established as serine and this change lasted until Perciformes. Tyrosine at the 5th position was infrequently changed into phenylalanine in Atheriniformes and Beloniformes and was observed in Perciformes as well as some Rodnetia of Mammalia.
In Perciformes, GnRH1 sequence experienced more diversification. The 2nd residue histidine was altered into leucine and the 5th tyrosine into phenylalanine as appeared in some animals, Atheriniformes, Atheriniformes, and Rodentia.
The 8th residue was distinctively modified into arginine in Scorpaeniformes, which is also observed in Siluriformes and Salmoniformes. The great diversity in the sequence of GnRH1 seems to have emerged from a transitional period into Anura. Similar to the primitive Vertebrata, Anura has asparagine as the 8th residue. Also it was converted into tryptophan. All these suggest that these extensive and diverse modifications to GnRH1 sequence were made in the time of Anura.
Reptilia has GnRH2 but not GnRH1. By the time Aves arised, the 8th residue was again altered into glutamate, which is uniquely featured in Aves only. In Mammalia, the 8th residue was arginine again and the resulting sequence was QHWSYGLRPG. This sequence, established in Diprotodontia, successfully got conserved in Rodentia, Scandentia, and Primates. Occasionally, other forms of GnRH such as QHWSFGLRPG or QYWSYGVRPG were seen in Rodentia. It can be deduced that such variations are due to the animals’ surrounding environments.
Comparing the sequence of GnRH1 to that of GnRH2 reveals that the homology is 70%. The discrepancies are at 3 positions - tyrosine to histidine at 5th, leucine to tryptophan at 7th, and arginine to tyrosine at 8th. Since the N- and C-termini are critical to receptor binding and activation signaling, these are fairly well conserved. Serine is fixed as the 4th residue in all forms of GnRH, which suggests that serine provides structural stability to the peptide.
GnRH2 first appeared in Chondrichthyes and stayed in most of Actinopterygii. Then it got passed down to Amphibia, Reptilia, some Aves, and some Mammalia. However, the fact that GnRH2 is only in some Aves (Columbiformes, Anseriformes, and Passeriformes) and Rodentia of Mammalia evokes some curiosity.
GnRH3 (QHWSYGWLPG) is also conserved but only appears in parts of Actinopterygii among Vertabrata (Mohamed et al., 2005; Mohamed & Khan, 2006). Unlike the typical GnRH1 found in Mammalia, the 7th and 8th residues are altered to tryptophan and leucine, respectively, resulting in 80% homology to GnRH1. Again, the N- and C-termini are well conserved. GnRH3 is not detected in Orders Chodrichthyes, primitive Actinopterygii, and some Actinopterygii. Perhaps the absence of GnRH3 is associated with sexual behaviors that GnRH3 regulates.
GnRH NUCLEIC ACID SEQUENCE ANALYSIS
Variations in the GnRH amino acid sequence described above are due to mutations on the nucleic acid sequence. The alterations result in modification of amino acid sequence expressed, regardless of whether the base substitutions are made by meiosis or external environmental factors. Substitutions can be made to any base at random. However, these substitutions do not drastically change the function of GnRH. Table 2 lists nucleic acid sequences of GnRH1.
The amino acids are abbreviated with single alphabets. The arrangement of Orders is within the evolutionary framework, with the most recent at top. Arabic numerals in the far-right column indicate number of variations in the sequence within the Order.
As mentioned above, both terminals of GnRH amino acid sequences are well conserved. However, there are some variations in the nucleic acid sequences. While the 1st and 2nd bases are inevitably preserved, the 3rd bases show great variations - synonymous substitution.
The GnRH-like peptide in Leptocardii of Cephalochordata (Roch et al., 2014) has 14 amino acids and is similar to GnRH2. Thus, there are no similarities in the nucleic acids of Cephalochordata GnRH-like peptide and vertebrate GnRH1. The nucleic acids in decapeptides detected in Phlebobranchia and Stolidobranchia in Tunicata do share some similarities.
The codon for pyroglutamate, the 1st residue in Leptocardii Cephalochordata GnRH-like peptide, is caa. This triple was continued to be passed down. Other nucleic acids in the peptide cannot be examined because the amino acids of the peptide are very different from those of GnRH1. Similarity to GnRH1 arose in the Ascidiacea, whose sequence has 10 residues as well. The first triplet in the Ascidiacea is cag or caa, which are also identified in the Cephalochordata. These two triplets were conserved from the Cephalochordata to Primates. The second triplet encoding histidine is cac in Stolidobranchia and cat in Phlebobrancchia. These triplets also got passed down as the 1st triplet, except for the Perciformes, whose can also be ctc. The third triplet tgg was conserved in all but the Petromyzontiformes, whose can be tac instead. The fourth residue is serine, encoded by tc* (* indicates any out of the 4 base choices).
The fifth residue can be varied depending on the Order. Most Actinopterygii, Amphibia, Aves, and Mammalia have tyrosine whereas Petromyzontiformes and some Actinopterygii have histidine instead. Additionally, some Vertebrata, Perciformes of Actinopterygii, and some Rodentia of Mammalia have phenylalanine. Lastly, Cypriniformes uniquely have arginine. Thus, the nucleic acids that encode the 5th residue vary widely. The initial base in the triplet is t in Stolidobranchia, which lasted through most Actinopterygii, Anura, Amphibia, Reptilia, Aves, and Mammalia. In Petromyzontiformes, which is the most primitive Vertebrata, the triplet begins with c, and this triplet is passed down to some Actinopterygii such as Clupeiformes, Cypriniformes, and Siluriformes. Lastly, in Phlebobranchia, it is switched to a, which results in asparagine (nonsynonymous substitution). Despite the diversity in the 5th residue, most have t or c as the initial base of the triplet, which evokes curiosity. The middle base is usually a or t, except for the Cypriniformes whose middle base is g. There are variations in the last base also, but they do not affect which amino acid gets encoded. Interesting to note is that ta- triplets would be a stop codon if the last base is a or g.
The 6th residue glycine is gg* in Vertebrata, which is also observed in Stolidobranchia Tunica. gg* is seen in all but the Petromyzontiformes, where it is changed to ga*. The 7th residue has some variations in its base triplet. From the Ascidiacea to Petromyzontiformes, the first base is t whereas from the Acipenseriformes to Primates, it is c. In Salmoniformes, the initial base is a, which results in encoding of methionine. The middle base is t or g in all cases. The 8th residue shows variations as well. Triplet att first appeared in Phlebobranchia and having a as the initial base was conserved up until Scorpaeniformes, which is just before Anura in the course of evolution. After Scorpaeniformes, the a was changed into c in almost all but Rodentia, Anura, and Phlebobranchia. Anura's cgg or tgg got modified to caa or cag in Aves, which encode all glutamine and contribute to the unique characteristics of Aves. The last two residues of the decapeptide sequence are well conserved, with the first two bases being cc and gg, respectively. The last base varies widely. All these variations in nucleic acids can explain the variations in amino acid sequence of GnRH1.
GnRH2 is described as ca* ca* tgg tc* ca* gg* tgg ta* cc* gg* and GnRH3 as ca* ca* tgg tc* ta* gg* tgg ct* cc* gg*. They share the commonality of having tgg as their 3rd and 7th triplets, which encode tryptophan. It may be these tryptophans that allow receptor binding and provide structural stability. Differences between GnRH2 and GnRH3 lie in the 1st base of 5th residue (c in GnRH2 and t in GnRH3) and first two bases of 8th residue (ta in GnRH2 and ct in GnRH3). These differences lead to encoding of different amino acids, which then contribute to differences in sexual behavior.
CONCLUSION
The analysis was performed by examining the amino acid and nucleic acid sequences of all GnRH identified in the Chordata from the evolutionary perspective. GnRH was first discovered in mammals as a decapeptide composed of 10 amino acids. As of present, there are 3 variations of GnRH - GnRH1, GnRH2, or GnRH3. They all share the N- and C-termini and glycine in the middle which allows for bending, and thus efficient binding to the receptor. The main function of GnRH1 is to produce germ cells and more extreme variations in the amino acid sequence are observed in the middle, at 5th, 7th, and 8th positions. Other substitutions in the sequence are perhaps due to variations in the nucleic acids, which were able to adapt to the environment. GnRH2 and GnRH3 are fairly well conserved, as they have the same amino acid sequence from Chondrichthyes to most Actinopterygii, Amphibia, Reptilia, some Aves, and Primates.
While there are great variations in the amino acids of GnRH1, its nucleic acids show some consistency. The nucleic acid sequence of GnRH1 is ca* ca* tgg tc* (t/c/a)a* gg* (t/c)(t/g)* (t/c/a)(t/g)* cc* gg*. The bases encoding 5th, 7th, and 8th residues are widely varied, which is consistent with the observation made for the amino acid sequence. Some changes in the base do not lead to the production of a different amino acid, and this is thought to maintain the function of GnRH1. GnRH2 has the sequence ca* ca* tgg tc* ca* gg* tgg ta* cc* gg*, which is conserved from Chondrichthyes to Primates. Again, it is possible that this conservation ensures the role of GnRH2 in energy balance and associative reproductive activity. Lastly, GnRH3 is ca* ca* tgg tc* ta* gg* tgg ct* cc* gg*. All three types of GnRH have tgg as its third triplet, which perhaps contributes to the structure and activity of GnRH.
Amino acids are the tools that allow for adaptation to the ambient environment and thus survival during evolutionary processes. Sequences that were chosen by natural selection have the capability to keep the species in the environment. Since variations of amino acids are due to alterations in nucleic acid sequences, it is safe to say that modifications made to nucleic acids determine the impact of evolutionary processes. Only the nucleic acids that increase the chances of survival get passed down while the others disappear. Investigating the changes that took place in both amino acid and nucleic acid sequences of GnRH reveals that GnRH has also been subjected to natural selection.